Genetically Predicted Levels of DNA Methylation Biomarkers and Breast Cancer Risk: Data From 228 951 Women of European Descent
- PMID: 31143935
- PMCID: PMC7073907
- DOI: 10.1093/jnci/djz109
Genetically Predicted Levels of DNA Methylation Biomarkers and Breast Cancer Risk: Data From 228 951 Women of European Descent
Abstract
Background: DNA methylation plays a critical role in breast cancer development. Previous studies have identified DNA methylation marks in white blood cells as promising biomarkers for breast cancer. However, these studies were limited by low statistical power and potential biases. Using a new methodology, we investigated DNA methylation marks for their associations with breast cancer risk.
Methods: Statistical models were built to predict levels of DNA methylation marks using genetic data and DNA methylation data from HumanMethylation450 BeadChip from the Framingham Heart Study (n = 1595). The prediction models were validated using data from the Women's Health Initiative (n = 883). We applied these models to genomewide association study (GWAS) data of 122 977 breast cancer patients and 105 974 controls to evaluate if the genetically predicted DNA methylation levels at CpG sites (CpGs) are associated with breast cancer risk. All statistical tests were two-sided.
Results: Of the 62 938 CpG sites CpGs investigated, statistically significant associations with breast cancer risk were observed for 450 CpGs at a Bonferroni-corrected threshold of P less than 7.94 × 10-7, including 45 CpGs residing in 18 genomic regions, that have not previously been associated with breast cancer risk. Of the remaining 405 CpGs located within 500 kilobase flaking regions of 70 GWAS-identified breast cancer risk variants, the associations for 11 CpGs were independent of GWAS-identified variants. Integrative analyses of genetic, DNA methylation, and gene expression data found that 38 CpGs may affect breast cancer risk through regulating expression of 21 genes.
Conclusion: Our new methodology can identify novel DNA methylation biomarkers for breast cancer risk and can be applied to other diseases.
© The Author(s) 2019. Published by Oxford University Press. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures
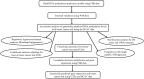


References
-
- DeSantis CE, Fedewa SA, Goding Sauer A, et al. Breast cancer statistics, 2015: convergence of incidence rates between black and white women. CA Cancer J Clin. 2016;66(1):31–42. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA148667/CA/NCI NIH HHS/United States
- C1287/A10710/CRUK_/Cancer Research UK/United Kingdom
- C1287/A16563/CRUK_/Cancer Research UK/United Kingdom
- C18281/A19169/CRUK_/Cancer Research UK/United Kingdom
- 16563/CRUK_/Cancer Research UK/United Kingdom
- 10118/CRUK_/Cancer Research UK/United Kingdom
- C1287/A10118/CRUK_/Cancer Research UK/United Kingdom
- U19 CA148065/CA/NCI NIH HHS/United States
- 10119/CRUK_/Cancer Research UK/United Kingdom
- 16561/CRUK_/Cancer Research UK/United Kingdom
- R01 CA158473/CA/NCI NIH HHS/United States
- 10124/CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Medical

