Neuronatin is a modifier of estrogen receptor-positive breast cancer incidence and outcome
- PMID: 31165373
- PMCID: PMC10317546
- DOI: 10.1007/s10549-019-05307-8
Neuronatin is a modifier of estrogen receptor-positive breast cancer incidence and outcome
Abstract
Purpose: Understanding the molecular mediators of breast cancer survival is critical for accurate disease prognosis and improving therapies. Here, we identified Neuronatin (NNAT) as a novel antiproliferative modifier of estrogen receptor-alpha (ER+) breast cancer.
Experimental design: Genomic regions harboring breast cancer modifiers were identified by congenic mapping in a rat model of carcinogen-induced mammary cancer. Tumors from susceptible and resistant congenics were analyzed by RNAseq to identify candidate genes. Candidates were prioritized by correlation with outcome, using a consensus of three breast cancer patient cohorts. NNAT was transgenically expressed in ER+ breast cancer lines (T47D and ZR75), followed by transcriptomic and phenotypic characterization.
Results: We identified a region on rat chromosome 3 (142-178 Mb) that modified mammary tumor incidence. RNAseq of the mammary tumors narrowed the candidate list to three differentially expressed genes: NNAT, SLC35C2, and FAM210B. NNAT mRNA and protein also correlated with survival in human breast cancer patients. Quantitative immunohistochemistry of NNAT protein revealed an inverse correlation with survival in a univariate analysis of patients with invasive ER+ breast cancer (training cohort: n = 444, HR = 0.62, p = 0.031; validation cohort: n = 430, HR = 0.48, p = 0.004). NNAT also held up as an independent predictor of survival after multivariable adjustment (HR = 0.64, p = 0.038). NNAT significantly reduced proliferation and migration of ER+ breast cancer cells, which coincided with altered expression of multiple related pathways.
Conclusions: Collectively, these data implicate NNAT as a novel mediator of cell proliferation and migration, which correlates with decreased tumorigenic potential and prolonged patient survival.
Keywords: Breast Cancer; Cell cycle; Estrogen; NNAT; Prognosis.
Conflict of interest statement
Figures



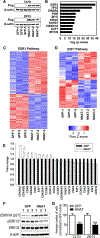


References
-
- Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clin 61(2):69–90 - PubMed
-
- Youlden DR, Cramb SM, Dunn NA, Muller JM, Pyke CM, Baade PD (2012) The descriptive epidemiology of female breast cancer: an international comparison of screening, incidence, survival and mortality. Cancer Epidemiol 36(3):237–248 - PubMed
-
- Stewart BW, Wild C, International Agency for Research on Cancer, World Health Organization (2014) World cancer report 2014. WHO Press, Lyon
-
- Pitale PM, Howse W, Gorbatyuk M (2017) Neuronatin protein in health and disease. J Cell Physiol 232(3):477–481 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

