Molecular Profiling and Functional Analysis of Macrophage-Derived Tumor Extracellular Vesicles
- PMID: 31167148
- PMCID: PMC6581796
- DOI: 10.1016/j.celrep.2019.05.008
Molecular Profiling and Functional Analysis of Macrophage-Derived Tumor Extracellular Vesicles
Abstract
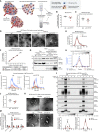
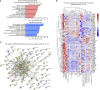
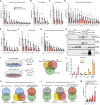
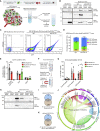
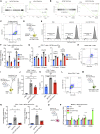
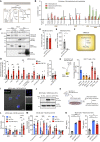
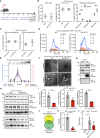
Extracellular vesicles (EVs), including exosomes, modulate multiple aspects of cancer biology. Tumor-associated macrophages (TAMs) secrete EVs, but their molecular features and functions are poorly characterized. Here, we report methodology for the enrichment, quantification, and proteomic and lipidomic analysis of EVs released from mouse TAMs (TAM-EVs). Compared to source TAMs, TAM-EVs present molecular profiles associated with a Th1/M1 polarization signature, enhanced inflammation and immune response, and a more favorable patient prognosis. Accordingly, enriched TAM-EV preparations promote T cell proliferation and activation ex vivo. TAM-EVs also contain bioactive lipids and biosynthetic enzymes, which may alter pro-inflammatory signaling in the cancer cells. Thus, whereas TAMs are largely immunosuppressive, their EVs may have the potential to stimulate, rather than limit, anti-tumor immunity.
Keywords: T cell response; exosome; extracellular vesicle; inflammation; lipid metabolism; lipidomics; proteomics; tumor microenvironment; tumor-associated macrophage.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Effect of colorectal cancer-derived extracellular vesicles on the immunophenotype and cytokine secretion profile of monocytes and macrophages.Cell Commun Signal. 2018 Apr 24;16(1):17. doi: 10.1186/s12964-018-0229-y. Cell Commun Signal. 2018. PMID: 29690889 Free PMC article.
-
A Comprehensive Proteomics Analysis Reveals a Secretory Path- and Status-Dependent Signature of Exosomes Released from Tumor-Associated Macrophages.J Proteome Res. 2015 Oct 2;14(10):4319-31. doi: 10.1021/acs.jproteome.5b00770. Epub 2015 Sep 8. J Proteome Res. 2015. PMID: 26312558
-
Extracellular vesicle-packaged miR-181c-5p from epithelial ovarian cancer cells promotes M2 polarization of tumor-associated macrophages via the KAT2B/HOXA10 axis.J Gene Med. 2022 Oct;24(10):e3446. doi: 10.1002/jgm.3446. Epub 2022 Sep 19. J Gene Med. 2022. PMID: 36027869
-
Roles of M1 Macrophages and Their Extracellular Vesicles in Cancer Therapy.Cells. 2024 Aug 26;13(17):1428. doi: 10.3390/cells13171428. Cells. 2024. PMID: 39273000 Free PMC article. Review.
-
Exploiting Manipulated Small Extracellular Vesicles to Subvert Immunosuppression at the Tumor Microenvironment through Mannose Receptor/CD206 Targeting.Int J Mol Sci. 2020 Aug 31;21(17):6318. doi: 10.3390/ijms21176318. Int J Mol Sci. 2020. PMID: 32878276 Free PMC article. Review.
Cited by
-
Functional and Therapeutic Significance of Tumor-Associated Macrophages in Colorectal Cancer.Front Oncol. 2022 Feb 2;12:781233. doi: 10.3389/fonc.2022.781233. eCollection 2022. Front Oncol. 2022. PMID: 35186730 Free PMC article. Review.
-
Anti-Cancer Role and Therapeutic Potential of Extracellular Vesicles.Cancers (Basel). 2021 Dec 15;13(24):6303. doi: 10.3390/cancers13246303. Cancers (Basel). 2021. PMID: 34944923 Free PMC article. Review.
-
Exosome-Mediated Crosstalk Between Tumor and Tumor-Associated Macrophages.Front Mol Biosci. 2021 Oct 13;8:764222. doi: 10.3389/fmolb.2021.764222. eCollection 2021. Front Mol Biosci. 2021. PMID: 34722637 Free PMC article. Review.
-
Osteocyte Apoptosis Contributes to Cold Exposure-induced Bone Loss.Front Bioeng Biotechnol. 2021 Nov 11;9:733582. doi: 10.3389/fbioe.2021.733582. eCollection 2021. Front Bioeng Biotechnol. 2021. PMID: 34858954 Free PMC article.
-
Metabolic Cancer-Macrophage Crosstalk in the Tumor Microenvironment.Biology (Basel). 2020 Nov 7;9(11):380. doi: 10.3390/biology9110380. Biology (Basel). 2020. PMID: 33171762 Free PMC article. Review.
References
-
- Baer C., Squadrito M.L., Laoui D., Thompson D., Hansen S.K., Kiialainen A., Hoves S., Ries C.H., Ooi C.H., De Palma M. Suppression of microRNA activity amplifies IFN-γ-induced macrophage activation and promotes anti-tumour immunity. Nat. Cell Biol. 2016;18:790–802. - PubMed
- Baer, C., Squadrito, M.L., Laoui, D., Thompson, D., Hansen, S.K., Kiialainen, A., Hoves, S., Ries, C.H., Ooi, C.H., and De Palma, M. (2016). Suppression of microRNA activity amplifies IFN-γ-induced macrophage activation and promotes anti-tumour immunity. Nat. Cell Biol. 18, 790-802. - PubMed
-
- Barreira da Silva R., Albert M.L. Mouse CD8+ T Cell Migration in vitro and CXCR3 Internalization Assays. Bio. Protoc. 2017;7:e2185. - PMC - PubMed
- Barreira da Silva, R., and Albert, M.L. (2017). Mouse CD8+ T Cell Migration in vitro and CXCR3 Internalization Assays. Bio. Protoc. 7, e2185. - PMC - PubMed
-
- Becker A., Thakur B.K., Weiss J.M., Kim H.S., Peinado H., Lyden D. Extracellular Vesicles in Cancer: Cell-to-Cell Mediators of Metastasis. Cancer Cell. 2016;30:836–848. - PMC - PubMed
- Becker, A., Thakur, B.K., Weiss, J.M., Kim, H.S., Peinado, H., and Lyden, D. (2016). Extracellular Vesicles in Cancer: Cell-to-Cell Mediators of Metastasis. Cancer Cell 30, 836-848. - PMC - PubMed
-
- Bobrie A., Colombo M., Krumeich S., Raposo G., Théry C. Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J. Extracell. Vesicles. 2012;1:1. - PMC - PubMed
- Bobrie, A., Colombo, M., Krumeich, S., Raposo, G., and Thery, C. (2012). Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J. Extracell. Vesicles 1, 1. - PMC - PubMed
-
- Boilard E. Extracellular vesicles and their content in bioactive lipid mediators: more than a sack of microRNA. J. Lipid Res. 2018;59:2037–2046. - PMC - PubMed
- Boilard, E. (2018). Extracellular vesicles and their content in bioactive lipid mediators: more than a sack of microRNA. J. Lipid Res. 59, 2037-2046. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

