Hepatitis B virus reverse transcriptase polymorphisms between treated and treatment-naïve chronically infected patients
- PMID: 31179360
- PMCID: PMC6531556
- DOI: 10.1007/s13337-018-00510-5
Hepatitis B virus reverse transcriptase polymorphisms between treated and treatment-naïve chronically infected patients
Abstract
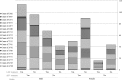
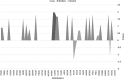
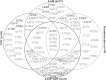
The aim of this study was investigation of variation(s) in the Hepatitis B virus (HBV) reverse transcriptase domain. 120 patients with chronic HBV infection recruited. 104 patients were received nucleos(t)ide analogs treatments. DNA extractions were done from plasma samples. Direct sequencing and alignment of Polymerase Chain Reaction products were applied for further analysis. HBV genotypes determined by NCBI's Genotyping Tool. Polymorphism(s) were detected by using DnaSP software. Of 120 samples, 98 were sequenced. All of products were HBV genotype D. 13/98 (13.27%) of patients had M539I/V substitutions corresponding to YMDD motif. FLLAQ to FLMAQ was observed among 22/98 (22.98) patients. Two substitutions N459Y and L515M were significantly correlated (R2 = 0.486 and R2 = 0.941 respectively) with FLLAQ motif variation. Mutation ratio among treatment-received patients to treatment-naïve patients was 0.2-0.6. Drug resistance conferring substitutions (DRCSs) were rtL180M (22/98), rtA194V (11/98), rtM204V (1/98), and rtM204I (11/98). Furthermore, six variants were observed among all patients. Appearance of DRCSs in HBV polymerase is a major obstacle to the virus treatments. In the present study, it was shown that DRCSs are more prevalent among treated patients. Therefore, replacement of current anti-viral regimen with novel anti-HBV drugs is warranted in the future.
Keywords: Chronic hepatitis B virus; Drug resistance conferring substitution; HBV reverse transcriptase; Hepatitis B virus polymerase; Nucleotide analogues resistance mutation.
Conflict of interest statement
Conflict of interestThe authors have no conflict of interest to declare.
Figures

Similar articles
-
Genotype-specific mutations in the polymerase gene of hepatitis B virus potentially associated with resistance to oral antiviral therapy.Antiviral Res. 2012 Dec;96(3):422-9. doi: 10.1016/j.antiviral.2012.09.014. Epub 2012 Sep 28. Antiviral Res. 2012. PMID: 23026293
-
Characterization of drug-resistance mutations in HBV D-genotype chronically infected patients, naïve to antiviral drugs.Antiviral Res. 2011 Nov;92(2):382-5. doi: 10.1016/j.antiviral.2011.08.013. Epub 2011 Sep 5. Antiviral Res. 2011. PMID: 21920388
-
The rtL229 substitutions in the reverse transcriptase region of hepatitis B virus (HBV) polymerase are potentially associated with lamivudine resistance as a compensatory mutation.J Clin Virol. 2012 May;54(1):66-72. doi: 10.1016/j.jcv.2012.02.003. Epub 2012 Mar 6. J Clin Virol. 2012. PMID: 22398037
-
Hepatitis B virus reverse transcriptase mutations in treatment Naïve chronic hepatitis B patients.J Med Virol. 2013 Jul;85(7):1155-62. doi: 10.1002/jmv.23608. J Med Virol. 2013. PMID: 23918533
-
Nucleos(t)ide analogues causes HBV S gene mutations and carcinogenesis.Hepatobiliary Pancreat Dis Int. 2016 Dec;15(6):579-586. doi: 10.1016/s1499-3872(16)60064-4. Hepatobiliary Pancreat Dis Int. 2016. PMID: 27919846 Review.
Cited by
-
A fragment-based drug discovery developed on ciclopirox for inhibition of Hepatitis B virus core protein: An in silico study.PLoS One. 2023 May 17;18(5):e0285941. doi: 10.1371/journal.pone.0285941. eCollection 2023. PLoS One. 2023. PMID: 37196004 Free PMC article.
-
The Antiviral Activity of Polyphenols.Mol Nutr Food Res. 2025 Aug;69(15):e70042. doi: 10.1002/mnfr.70042. Epub 2025 Apr 1. Mol Nutr Food Res. 2025. PMID: 40166854 Free PMC article. Review.
-
Pharmacophore modeling and QSAR analysis of anti-HBV flavonols.PLoS One. 2025 Jan 13;20(1):e0316765. doi: 10.1371/journal.pone.0316765. eCollection 2025. PLoS One. 2025. PMID: 39804828 Free PMC article.
-
An overview of anti-Hepatitis B virus flavonoids and their mechanisms of action.Front Cell Infect Microbiol. 2024 Feb 29;14:1356003. doi: 10.3389/fcimb.2024.1356003. eCollection 2024. Front Cell Infect Microbiol. 2024. PMID: 38487354 Free PMC article. Review.
References
-
- Jardi R, Rodriguez-Frias F, Schaper M, Ruiz G, Elefsiniotis I, Esteban R, et al. Hepatitis B virus polymerase variants associated with entecavir drug resistance in treatment-naive patients. J Viral Hepat. 2007;14:835–840. - PubMed
LinkOut - more resources
Full Text Sources

