DNA Methylation Changes in the Sperm of Captive-Reared Fish: A Route to Epigenetic Introgression in Wild Populations
- PMID: 31180510
- PMCID: PMC6759066
- DOI: 10.1093/molbev/msz135
DNA Methylation Changes in the Sperm of Captive-Reared Fish: A Route to Epigenetic Introgression in Wild Populations
Abstract
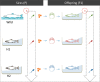
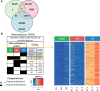
Interbreeding between hatchery-reared and wild fish, through deliberate stocking or escapes from fish farms, can result in rapid phenotypic and gene expression changes in hybrids, but the underlying mechanisms are unknown. We assessed if one generation of captive breeding was sufficient to generate inter- and/or transgenerational epigenetic modifications in Atlantic salmon. We found that the sperm of wild and captive-reared males differed in methylated regions consistent with early epigenetic signatures of domestication. Some of the epigenetic marks that differed between hatchery and wild males affected genes related to transcription, neural development, olfaction, and aggression, and were maintained in the offspring beyond developmental reprogramming. Our findings suggest that rearing in captivity may trigger epigenetic modifications in the sperm of hatchery fish that could explain the rapid phenotypic and genetic changes observed among hybrid fish. Epigenetic introgression via fish sperm represents a previously unappreciated mechanism that could compromise locally adapted fish populations.
Keywords: Salmo salar; epigenetic inheritance; DNA methylation; domestication.
© The Author(s) 2018. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures

Similar articles
-
Captive rearing effects on the methylome of Atlantic salmon after oceanic migration: Sex-specificity and intergenerational stability.Mol Ecol Resour. 2025 Jul;25(5):e13766. doi: 10.1111/1755-0998.13766. Epub 2023 Feb 23. Mol Ecol Resour. 2025. PMID: 36760032 Free PMC article.
-
Environment-driven reprogramming of gamete DNA methylation occurs during maturation and is transmitted intergenerationally in Atlantic Salmon.G3 (Bethesda). 2021 Dec 8;11(12):jkab353. doi: 10.1093/g3journal/jkab353. G3 (Bethesda). 2021. PMID: 34849830 Free PMC article.
-
Characterization of Genetic and Epigenetic Variation in Sperm and Red Blood Cells from Adult Hatchery and Natural-Origin Steelhead, Oncorhynchus mykiss.G3 (Bethesda). 2018 Nov 6;8(11):3723-3736. doi: 10.1534/g3.118.200458. G3 (Bethesda). 2018. PMID: 30275172 Free PMC article.
-
On the reproductive success of early-generation hatchery fish in the wild.Evol Appl. 2014 Sep;7(8):883-96. doi: 10.1111/eva.12183. Epub 2014 Jul 9. Evol Appl. 2014. PMID: 25469167 Free PMC article. Review.
-
Transgenerational epigenetic inheritance in mammals: how good is the evidence?FASEB J. 2016 Jul;30(7):2457-65. doi: 10.1096/fj.201500083. Epub 2016 Apr 1. FASEB J. 2016. PMID: 27037350 Review.
Cited by
-
Captive rearing effects on the methylome of Atlantic salmon after oceanic migration: Sex-specificity and intergenerational stability.Mol Ecol Resour. 2025 Jul;25(5):e13766. doi: 10.1111/1755-0998.13766. Epub 2023 Feb 23. Mol Ecol Resour. 2025. PMID: 36760032 Free PMC article.
-
The evolutionary consequences of interactions between the epigenome, the genome and the environment.Evol Appl. 2024 Jul 23;17(7):e13730. doi: 10.1111/eva.13730. eCollection 2024 Jul. Evol Appl. 2024. PMID: 39050763 Free PMC article. Review.
-
A new approach to decode DNA methylome and genomic variants simultaneously from double strand bisulfite sequencing.Brief Bioinform. 2021 Nov 5;22(6):bbab201. doi: 10.1093/bib/bbab201. Brief Bioinform. 2021. PMID: 34058751 Free PMC article.
-
Hatchery supplementation provides a demographic boost but alters age composition of sockeye salmon in Auke Lake, Southeast Alaska.Evol Appl. 2024 Feb 7;17(2):e13640. doi: 10.1111/eva.13640. eCollection 2024 Feb. Evol Appl. 2024. PMID: 38333553 Free PMC article.
-
Epigenomic modifications induced by hatchery rearing persist in germ line cells of adult salmon after their oceanic migration.Evol Appl. 2021 May 4;14(10):2402-2413. doi: 10.1111/eva.13235. eCollection 2021 Oct. Evol Appl. 2021. PMID: 34745334 Free PMC article.
References
-
- Andrews S. 2015. A tool to visualise and analyse high throughput mapped sequence data. Available from: http://www. bioinformatics. babraham. ac. uk/projects/seqmonk. Last accessed on March 20, 2018.
-
- Araki H, Cooper B, Blouin MS.. 2007. Genetic effects of captive breeding cause a rapid, cumulative fitness decline in the wild. Science 3185847:100–103. - PubMed
-
- Araki H, Schmid C.. 2010. Is hatchery stocking a help or harm? Evidence, limitations and future directions in ecological and genetic surveys. Aquaculture 308:S2–S11.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

