A Candidate for Multitopic Probes for Ligand Discovery in Dynamic Combinatorial Chemistry
- PMID: 31181809
- PMCID: PMC6600254
- DOI: 10.3390/molecules24112166
A Candidate for Multitopic Probes for Ligand Discovery in Dynamic Combinatorial Chemistry
Abstract
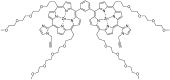
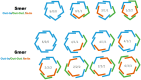
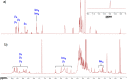
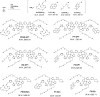
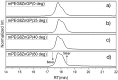
Multifunctionalized materials are expected to be versatile probes to find specific interactions between a ligand and a target biomaterial. Thus, efficient methods to prepare possible combinations of the functionalities is desired. The concept of dynamic combinatorial chemistry (DCC) is ideal for the generation of any possible combination, as well as screening for target biomaterials. Here, we propose a new molecular design of multitopic probes for ligand discovery in DCC. We synthesized a new Gable Porphyrin, GP1, having prop-2-yne groups as a scaffold to introduce various functional groups. GP1 is a bis(imidazolylporphyrinatozinc) compound connected through a 1,3-phenylene moiety, and it gives macrocycles spontaneously and quantitatively by strong imidazole-to-zinc complementary coordination. Some different types of functional groups were introduced into GP1 in high yields. Formation of heterogeneous macrocycles composed of GP1 derivatives having different types of substituents was accomplished under equilibrium conditions. These results promise that enormous numbers of macrocycles having various functional groups can be provided when the kinds of GP components increase. These features are desirable for DCC, and the present system using GP1 is a potential candidate to provide a dynamic combinatorial library of multitopic probes to discover specific interactions between a ligand and a biomaterial.
Keywords: amphiphilic; complementary coordination; copper(I)-catalyzed alkyne-azide cycloaddition (CuAAC); dynamic combinatorial chemistry (DCC); dynamic combinatorial library (DCL); gel permeation chromatography (GPC); multifunctionalized material; supramolecular macrocycle; zinc porphyrin.
Conflict of interest statement
The authors declare no conflict of interest.
Figures








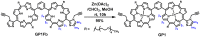











References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

