IFN-I and IL-22 mediate protective effects of intestinal viral infection
- PMID: 31182797
- PMCID: PMC6871771
- DOI: 10.1038/s41564-019-0470-1
IFN-I and IL-22 mediate protective effects of intestinal viral infection
Abstract
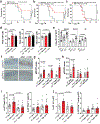
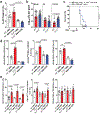
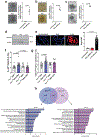
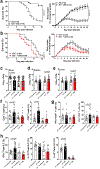
Products derived from bacterial members of the gut microbiota evoke immune signalling pathways of the host that promote immunity and barrier function in the intestine. How immune reactions to enteric viruses support intestinal homeostasis is unknown. We recently demonstrated that infection by murine norovirus (MNV) reverses intestinal abnormalities following depletion of bacteria, indicating that an intestinal animal virus can provide cues to the host that are typically attributed to the microbiota. Here, we elucidate mechanisms by which MNV evokes protective responses from the host. We identify an important role for the viral protein NS1/2 in establishing local replication and a type I interferon (IFN-I) response in the colon. We further show that IFN-I acts on intestinal epithelial cells to increase the proportion of CCR2-dependent macrophages and interleukin (IL)-22-producing innate lymphoid cells, which in turn promote pSTAT3 signalling in intestinal epithelial cells and protection from intestinal injury. In addition, we demonstrate that MNV provides a striking IL-22-dependent protection against early-life lethal infection by Citrobacter rodentium. These findings demonstrate novel ways in which a viral member of the microbiota fortifies the intestinal barrier during chemical injury and infectious challenges.
Conflict of interest statement
Competing Financial Interests
K.C. has consulted for PureTech Health and AbbVie Inc. and is an inventor on U.S. patent application 62/608,404.
Figures



References
-
- Basic M, et al. Norovirus triggered microbiota-driven mucosal inflammation in interleukin 10-deficient mice. Inflamm Bowel Dis 20, 431–443 (2014). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

