Stepwise Development of an in vitro Continuous Fermentation Model for the Murine Caecal Microbiota
- PMID: 31191488
- PMCID: PMC6548829
- DOI: 10.3389/fmicb.2019.01166
Stepwise Development of an in vitro Continuous Fermentation Model for the Murine Caecal Microbiota
Abstract
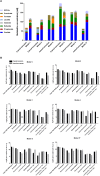
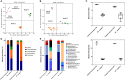
Murine models are valuable tools to study the role of gut microbiota in health or disease. However, murine and human microbiota differ in species composition, so further investigation of the murine gut microbiota is important to gain a better mechanistic understanding. Continuous in vitro fermentation models are powerful tools to investigate microbe-microbe interactions while circumventing animal testing and host confounding factors, but are lacking for murine gut microbiota. We therefore developed a novel continuous fermentation model based on the PolyFermS platform adapted to the murine caecum and inoculated with immobilized caecal microbiota. We followed a stepwise model development approach by adjusting parameters [pH, retention time (RT), growth medium] to reach fermentation metabolite profiles and marker bacterial levels similar to the inoculum. The final model had a stable and inoculum-alike fermentation profile during continuous operation. A lower pH during startup and continuous operation stimulated bacterial fermentation (115 mM short-chain fatty acids at pH 7 to 159 mM at pH 6.5). Adjustments to nutritive medium, a decreased pH and increased RT helped control the in vitro Enterobacteriaceae levels, which often bloom in fermentation models, to 6.6 log gene copies/mL in final model. In parallel, the Lactobacillus, Lachnospiraceae, and Ruminococcaceae levels were better maintained in vitro with concentrations of 8.5 log gene copies/mL, 8.8 log gene copies/mL and 7.5 log gene copies/mL, respectively, in the final model. An independent repetition with final model parameters showed reproducible results in maintaining the inoculum fermentation metabolite profile and its marker bacterial levels. Microbiota community analysis of the final model showed a decreased bacterial diversity and compositional differences compared to caecal inoculum microbiota. Most of the caecal bacterial families were represented in vitro, but taxa of the Muribaculaceae family were not maintained. Functional metagenomics prediction showed conserved metabolic and functional KEGG pathways between in vitro and caecal inoculum microbiota. To conclude, we showed that a rational and stepwise approach allowed us to model in vitro the murine caecal microbiota and functions. Our model is a first step to develop murine microbiota model systems and offers the potential to study microbiota functionality and structure ex vivo.
Keywords: C57BL/6; cultivation; in vitro model; microbiome; mouse caecum.
Figures

Similar articles
-
In vitro Modeling of Chicken Cecal Microbiota Ecology and Metabolism Using the PolyFermS Platform.Front Microbiol. 2021 Dec 20;12:780092. doi: 10.3389/fmicb.2021.780092. eCollection 2021. Front Microbiol. 2021. PMID: 34987487 Free PMC article.
-
Long-term continuous cultivation of Kenyan infant fecal microbiota using the host adapted PolyFermS model.Res Sq [Preprint]. 2023 Jul 7:rs.3.rs-3101157. doi: 10.21203/rs.3.rs-3101157/v1. Res Sq. 2023. Update in: Sci Rep. 2023 Nov 23;13(1):20563. doi: 10.1038/s41598-023-47131-7. PMID: 37461546 Free PMC article. Updated. Preprint.
-
Design and Investigation of PolyFermS In Vitro Continuous Fermentation Models Inoculated with Immobilized Fecal Microbiota Mimicking the Elderly Colon.PLoS One. 2015 Nov 11;10(11):e0142793. doi: 10.1371/journal.pone.0142793. eCollection 2015. PLoS One. 2015. PMID: 26559530 Free PMC article.
-
Advances in in vitro cultivation techniques for comprehensive analysis of human gut microbiome.Biotechnol Adv. 2025 Sep;82:108595. doi: 10.1016/j.biotechadv.2025.108595. Epub 2025 May 13. Biotechnol Adv. 2025. PMID: 40374084 Review.
-
Application of in vitro gut fermentation models to food components: A review.Food Sci Biotechnol. 2016 Mar 31;25(Suppl 1):1-7. doi: 10.1007/s10068-016-0091-x. eCollection 2016. Food Sci Biotechnol. 2016. PMID: 30263479 Free PMC article. Review.
Cited by
-
Yeast Fermentate-Mediated Reduction of Salmonella Reading and Typhimurium in an in vitro Turkey Cecal Culture Model.Front Microbiol. 2021 Apr 15;12:645301. doi: 10.3389/fmicb.2021.645301. eCollection 2021. Front Microbiol. 2021. PMID: 33936004 Free PMC article.
-
Modeling Dynamics of Human Gut Microbiota Derived from Gluten Metabolism: Obtention, Maintenance and Characterization of Complex Microbial Communities.Int J Mol Sci. 2024 Apr 4;25(7):4013. doi: 10.3390/ijms25074013. Int J Mol Sci. 2024. PMID: 38612823 Free PMC article.
-
GABA Production by Human Intestinal Bacteroides spp.: Prevalence, Regulation, and Role in Acid Stress Tolerance.Front Microbiol. 2021 Apr 15;12:656895. doi: 10.3389/fmicb.2021.656895. eCollection 2021. Front Microbiol. 2021. PMID: 33936013 Free PMC article.
-
In vitro Modeling of Chicken Cecal Microbiota Ecology and Metabolism Using the PolyFermS Platform.Front Microbiol. 2021 Dec 20;12:780092. doi: 10.3389/fmicb.2021.780092. eCollection 2021. Front Microbiol. 2021. PMID: 34987487 Free PMC article.
-
New Horizons in Probiotics: Unraveling the Potential of Edible Microbial Polysaccharides through In Vitro Digestion Models.Foods. 2024 Feb 26;13(5):713. doi: 10.3390/foods13050713. Foods. 2024. PMID: 38472826 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

