Community Structure of Bacteria Associated With Drifting Sargassum horneri, the Causative Species of Golden Tide in the Yellow Sea
- PMID: 31191503
- PMCID: PMC6546727
- DOI: 10.3389/fmicb.2019.01192
Community Structure of Bacteria Associated With Drifting Sargassum horneri, the Causative Species of Golden Tide in the Yellow Sea
Abstract
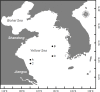
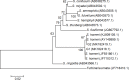
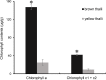
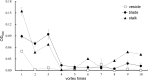
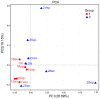
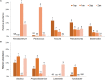
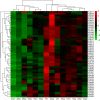
Golden tides dominated by Sargassum spp. are occurring at an accelerated rate worldwide. In China, Sargassum has started to bloom in the Yellow Sea and led to tremendous economic losses, but the underlying biological causes and mechanisms are still unclear. Although algae-associated bacteria were suggested to play crucial roles in algal blooms, the profiles of bacteria associated with drifting Sargassum remain unexplored. In this study, the community structures and functions of Sargassum-associated bacteria were analyzed using the high-throughput sequencing data of the V5-V7 hypervariable region of the 16S rRNA gene. Molecular identification revealed that the golden tide analyzed in the Yellow Sea was dominated by a single species, Sargassum horneri. They were a healthy brown color nearshore but were yellow offshore with significantly decreased chlorophyll contents (P < 0.01), which indicates that yellow S. horneri was under physiological stress. The structural and functional analyses of bacterial communities indicated that the drifting S. horneri had an obvious selectivity on their associated bacteria against surrounding seawater. Although the bacterial communities phylogenetically differed between brown and yellow S. horneri (P < 0.01), their dominant functions were all nitrogen and iron transporters, which strongly indicates microbial contribution to blooming of the algal host. For the first time, potential epiphytic and endophytic bacteria associated with Sargassum were independently analyzed by a modified co-vortex method with silica sand. We showed that the composition of dominant endophytes, mainly Bacillus and Propionibacterium, was relatively consistent regardless of host status, whereas the epiphytic operational taxonomic units (OTUs) greatly varied in response to weakness of host status; however, dominant functions were consistent at elevated intensities, which might protect the host from stress related to nitrogen or iron deficiency. Thus, we propose that host physiological status at different intensities of functional demands, which were related to variable environmental conditions, may be a critical factor that influences the assembly of epiphytic bacterial communities. This study provided new insight into the structure and potential functions of associated bacteria with golden tide blooms.
Keywords: 16S rRNA gene; Sargassum horneri; bacterial community; golden tide; high-throughput sequencing; the Yellow Sea.
Figures





Similar articles
-
Effects of increased CO2 and temperature on the physiological characteristics of the golden tide blooming macroalgae Sargassum horneri in the Yellow Sea, China.Mar Pollut Bull. 2019 Sep;146:639-644. doi: 10.1016/j.marpolbul.2019.07.025. Epub 2019 Jul 17. Mar Pollut Bull. 2019. PMID: 31426203
-
Interannual variations of Sargassum blooms in the Yellow Sea and East China Sea during 2017-2021.Harmful Algae. 2023 Jul;126:102451. doi: 10.1016/j.hal.2023.102451. Epub 2023 May 6. Harmful Algae. 2023. PMID: 37290886
-
An anomalous bi-macroalgal bloom caused by Ulva and Sargassum seaweeds during spring to summer of 2017 in the western Yellow Sea, China.Harmful Algae. 2020 Mar;93:101760. doi: 10.1016/j.hal.2020.101760. Epub 2020 Feb 28. Harmful Algae. 2020. PMID: 32307078
-
Harmful macroalgal blooms (HMBs) in China's coastal water: Green and golden tides.Harmful Algae. 2021 Jul;107:102061. doi: 10.1016/j.hal.2021.102061. Epub 2021 Jun 13. Harmful Algae. 2021. PMID: 34456020 Review.
-
Where does floating Sargassum in the East China Sea come from?Harmful Algae. 2023 Nov;129:102523. doi: 10.1016/j.hal.2023.102523. Epub 2023 Oct 10. Harmful Algae. 2023. PMID: 37951622 Review.
Cited by
-
Halotolerant Bacteria from Genus Nesterenkonia sandarakina VSA9 as a Potential Polyhydroxyalkanoate Producer.Curr Microbiol. 2024 Jan 3;81(1):53. doi: 10.1007/s00284-023-03569-6. Curr Microbiol. 2024. PMID: 38172411
-
Easy Removal of Epiphytic Bacteria on Ulva (Ulvophyceae, Chlorophyta) by Vortex with Silica Sands.Microorganisms. 2022 Feb 21;10(2):476. doi: 10.3390/microorganisms10020476. Microorganisms. 2022. PMID: 35208930 Free PMC article.
-
Bacillus velezensis TB918 mitigates garlic dry rot disease by forming consortia with Pseudomonas in the rhizosphere and bulb.Front Microbiol. 2025 Apr 15;16:1567108. doi: 10.3389/fmicb.2025.1567108. eCollection 2025. Front Microbiol. 2025. PMID: 40303477 Free PMC article.
-
Composition and Functional Diversity of Epiphytic Bacterial and Fungal Communities on Marine Macrophytes in an Intertidal Zone.Front Microbiol. 2022 Mar 18;13:839465. doi: 10.3389/fmicb.2022.839465. eCollection 2022. Front Microbiol. 2022. PMID: 35369473 Free PMC article.
-
Epibiotic Bacteria Isolated from the Non-Indigenous Species Codium fragile ssp. fragile: Identification, Characterization, and Biotechnological Potential.Microorganisms. 2024 Aug 30;12(9):1803. doi: 10.3390/microorganisms12091803. Microorganisms. 2024. PMID: 39338477 Free PMC article.
References
-
- Ahmed E. F., Hassan H. M., Rateb M. E., Abdel-Wahab N., Sameer S., Aly Taie H. A., et al. (2016). A comparative biochemical study on two marine endophytes, bacterium SRCnm and Bacillus sp. JS, isolated from red sea algae. Pak. J. Pharm. Sci. 29 17–26. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

