F8 Inversions at Xq28 Causing Hemophilia A Are Associated With Specific Methylation Changes: Implication for Molecular Epigenetic Diagnosis
- PMID: 31191618
- PMCID: PMC6548806
- DOI: 10.3389/fgene.2019.00508
F8 Inversions at Xq28 Causing Hemophilia A Are Associated With Specific Methylation Changes: Implication for Molecular Epigenetic Diagnosis
Abstract
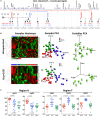
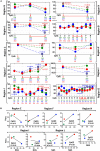
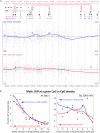
Diverse DNA structural variations (SVs) in human cancers and several other diseases are well documented. For genomic inversions in particular, the disease causing mechanism may not be clear, especially if the inversion border does not cross a coding sequence. Understanding about the molecular processes of these inverted genomic sequences, in a mainly epigenetic context, may provide additional information regarding sequence-specific regulation of gene expression in human diseases. Herein, we study one such inversion hotspot at Xq28, which leads to the disruption of F8 gene and results in hemophilia A phenotype. To determine the epigenetic consequence of this rearrangement, we evaluated DNA methylation levels of 12 CpG rich regions with the coverage of 550 kb by using bisulfite-pyrosequencing and next-generation sequencing (NGS)-based bisulfite re-sequencing enrichment assay. Our results show that this inversion prone area harbors widespread methylation changes at the studied regions. However, only 5/12 regions showed significant methylation changes, specifically in case of intron 1 inversion (two regions), intron 22 inversion (two regions) and one common region in both inversions. Interestingly, these aberrant methylated regions were found to be overlapping with the inversion proximities. In addition, two CpG sites reached 100% sensitivity and specificity to discriminate wild type from intron 22 and intron 1 inversion samples. While we found age to be an influencing factor on methylation levels at some regions, covariate analysis still confirms the differential methylation induced by inversion, regardless of age. The hemophilia A methylation inversion "HAMI" assay provides an advantage over conventional PCR-based methods, which may not detect novel rare genomic rearrangements. Taken together, we showed that genomic inversions in the F8 (Xq28) region are associated with detectable changes in methylation levels and can be used as an epigenetic diagnostic marker.
Keywords: DNA methylation; epigenetic; hemophilia; inversion; molecular diagnosis; structural rearrangement.
Figures

References
-
- Andrikovics H., Klein I., Bors A., Nemes L., Marosi A., Varadi A., et al. (2003). Analysis of large structural changes of the factor VIII gene, involving intron 1 and 22, in severe hemophilia A. Haematologica 88 778–784. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

