Social status regulates the hepatic miRNAome in rainbow trout: Implications for posttranscriptional regulation of metabolic pathways
- PMID: 31194802
- PMCID: PMC6563994
- DOI: 10.1371/journal.pone.0217978
Social status regulates the hepatic miRNAome in rainbow trout: Implications for posttranscriptional regulation of metabolic pathways
Erratum in
-
Correction: Social status regulates the hepatic miRNAome in rainbow trout: Implications for posttranscriptional regulation of metabolic pathways.PLoS One. 2020 May 21;15(5):e0233827. doi: 10.1371/journal.pone.0233827. eCollection 2020. PLoS One. 2020. PMID: 32437451 Free PMC article.
Abstract
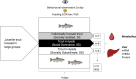
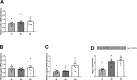
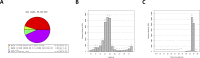
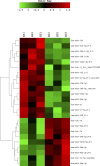
Juvenile rainbow trout develop social hierarchies when held in dyads, and the development of socially subordinate (SS) and social dominance (SD) phenotypes in this context has been linked to specific changes in the hepatic energy metabolism of all major macronutrients. Following our recently reported finding that transcript abundance of drosha, a key component of the microRNA (miRNA) biogenesis pathway, is increased in paired juvenile rainbow trout irrespective of social status compared to socially isolated (SI) controls, we here determined global changes of the hepatic miRNA pathway genes in detail at the transcript and protein level. Both SD and SS rainbow trout exhibited increased Ago2 protein abundance compared to SI rainbow trout, suggesting that hepatic miRNA function is increased in rainbow trout maintained in dyads. Given the well-described differences in hepatic intermediary metabolism between SD and SS rainbow trout, and the important role of miRNAs in the posttranscriptional regulation of metabolic pathways, we also identified changes in hepatic miRNA abundance between SS and SD rainbow trout using small RNA next generation sequencing. We identified a total of 24 differentially regulated miRNAs, with 15 miRNAs that exhibited increased expression, and 9 miRNAs that exhibited decreased expression in the liver of SS trout compared to SD trout. To identify potential miRNA-dependent posttranscriptional regulatory pathways important for social status-dependent regulation of hepatic metabolism in rainbow trout, we used an in silico miRNA target prediction and pathway enrichment approach. We identified enrichment for pathways related to metabolism of carbohydrates, lipids and proteins in addition to organelle-specific processes involved in energy metabolism, especially mitochondrial fusion and fission. Select predicted miRNA-mRNA target pairs within these categories were quantitatively analyzed by real-time RT-PCR to validate candidates for future studies that will probe the functional metabolic roles of specific hepatic miRNAs in the development of SD and SS metabolic phenotypes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Profiling the rainbow trout hepatic miRNAome under diet-induced hyperglycemia.Physiol Genomics. 2019 Sep 1;51(9):411-431. doi: 10.1152/physiolgenomics.00032.2019. Epub 2019 Jul 8. Physiol Genomics. 2019. PMID: 31282806 Free PMC article.
-
Meta-analysis of differentially-regulated hepatic microRNAs identifies candidate post-transcriptional regulation networks of intermediary metabolism in rainbow trout.Comp Biochem Physiol Part D Genomics Proteomics. 2020 Dec;36:100750. doi: 10.1016/j.cbd.2020.100750. Epub 2020 Oct 3. Comp Biochem Physiol Part D Genomics Proteomics. 2020. PMID: 33038710
-
Postprandial regulation of hepatic microRNAs predicted to target the insulin pathway in rainbow trout.PLoS One. 2012;7(6):e38604. doi: 10.1371/journal.pone.0038604. Epub 2012 Jun 6. PLoS One. 2012. PMID: 22701681 Free PMC article.
-
Micromanaging metabolism-a role for miRNAs in teleost energy metabolism.Comp Biochem Physiol B Biochem Mol Biol. 2016 Sep;199:115-125. doi: 10.1016/j.cbpb.2015.09.001. Epub 2015 Sep 15. Comp Biochem Physiol B Biochem Mol Biol. 2016. PMID: 26384523 Review.
-
Energizing miRNA research: a review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways.Mol Genet Metab. 2007 Jul;91(3):209-17. doi: 10.1016/j.ymgme.2007.03.011. Epub 2007 May 22. Mol Genet Metab. 2007. PMID: 17521938 Free PMC article. Review.
Cited by
-
High Throughput Sequencing of MicroRNA in Rainbow Trout Plasma, Mucus, and Surrounding Water Following Acute Stress.Front Physiol. 2021 Jan 13;11:588313. doi: 10.3389/fphys.2020.588313. eCollection 2020. Front Physiol. 2021. PMID: 33519501 Free PMC article.
-
Chronic social stress alters protein metabolism in juvenile rainbow trout, Oncorhynchus mykiss.J Comp Physiol B. 2021 May;191(3):517-530. doi: 10.1007/s00360-021-01340-6. Epub 2021 Mar 12. J Comp Physiol B. 2021. PMID: 33712903 Free PMC article.
-
Methylene blue at recommended concentrations alters metabolism in early zebrafish development.Commun Biol. 2025 Jan 25;8(1):120. doi: 10.1038/s42003-025-07471-8. Commun Biol. 2025. PMID: 39856203 Free PMC article.
-
Correction: Social status regulates the hepatic miRNAome in rainbow trout: Implications for posttranscriptional regulation of metabolic pathways.PLoS One. 2020 May 21;15(5):e0233827. doi: 10.1371/journal.pone.0233827. eCollection 2020. PLoS One. 2020. PMID: 32437451 Free PMC article.
References
-
- Abbott JC, Dill LM. Patterns of Aggressive Attack in Juvenile Steelhead Trout (Salmo gairdneri). Can J Fish Aquat Sci. 1985. November 1;42(11):1702–6.
-
- Elliott JM. Mechanisms Responsible for Population Regulation in Young Migratory Trout, Salmo trutta. III. The Role of Territorial Behaviour. Journal of Animal Ecology. 1990;59(3):803–18.
-
- Metcalfe NB. Intraspecific variation in competitive ability and food intake in salmonids: consequences for energy budgets and growth rates J Fish Biol. 1986. May 28: 525–531.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

