Trouble is coming: Signaling pathways that regulate general stress responses in bacteria
- PMID: 31197038
- PMCID: PMC6682744
- DOI: 10.1074/jbc.REV119.005593
Trouble is coming: Signaling pathways that regulate general stress responses in bacteria
Abstract
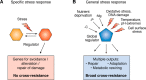
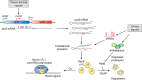
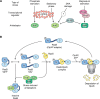
Bacteria can rapidly and reversibly respond to changing environments via complex transcriptional and post-transcriptional regulatory mechanisms. Many of these adaptations are specific, with the regulatory output tailored to the inducing signal (for instance, repairing damage to cell components or improving acquisition and use of growth-limiting nutrients). However, the general stress response, activated in bacterial cells entering stationary phase or subjected to nutrient depletion or cellular damage, is unique in that its common, broad output is induced in response to many different signals. In many different bacteria, the key regulator for the general stress response is a specialized sigma factor, the promoter specificity subunit of RNA polymerase. The availability or activity of the sigma factor is regulated by complex regulatory circuits, the majority of which are post-transcriptional. In Escherichia coli, multiple small regulatory RNAs, each made in response to a different signal, positively regulate translation of the general stress response sigma factor RpoS. Stability of RpoS is regulated by multiple anti-adaptor proteins that are also synthesized in response to different signals. In this review, the modes of signaling to and levels of regulation of the E. coli general stress response are discussed. They are also used as a basis for comparison with the general stress response in other bacteria with the aim of extracting key principles that are common among different species and highlighting important unanswered questions.
Keywords: ClpXP; Escherichia coli (E. coli); Hfq protein; RNA; RNA polymerase; RpoS; anti-adaptor; general stress response; prokaryotic signal-transduction; proteolysis.
Conflict of interest statement
The author declares that she has no conflicts of interest with the contents of this article
Figures


References
-
- Hamdallah I., Torok N., Bischof K. M., Majdalani N., Chadalavada S., Mdluli N., Creamer K. E., Clark M., Holdener C., Basting P. J., Gottesman S., and Slonczewski J. L. (2018) Experimental evolution of Escherichia coli K-12 at high pH and with RpoS induction. Appl. Environ. Microbiol. 84, e00520–18 10.1128/AEM.00520-18 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

