The yeast scavenger decapping enzyme DcpS and its application for in vitro RNA recapping
- PMID: 31197197
- PMCID: PMC6565619
- DOI: 10.1038/s41598-019-45083-5
The yeast scavenger decapping enzyme DcpS and its application for in vitro RNA recapping
Abstract
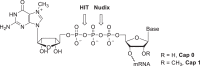
Eukaryotic mRNAs are modified at their 5' end early during transcription by the addition of N7-methylguanosine (m7G), which forms the "cap" on the first 5' nucleotide. Identification of the 5' nucleotide on mRNA is necessary for determination of the Transcription Start Site (TSS). We explored the effect of various reaction conditions on the activity of the yeast scavenger mRNA decapping enzyme DcpS and examined decapping of 30 chemically distinct cap structures varying the state of methylation, sugar, phosphate linkage, and base composition on 25mer RNA oligonucleotides. Contrary to the generally accepted belief that DcpS enzymes only decap short oligonucleotides, we found that the yeast scavenger decapping enzyme decaps RNA transcripts as long as 1400 nucleotides. Further, we validated the application of yDcpS for enriching capped RNA using a strategy of specifically tagging the 5' end of capped RNA by first decapping and then recapping it with an affinity-tagged guanosine nucleotide.
Conflict of interest statement
All authors are employees of New England Biolabs. New England Biolabs commercializes reagents for molecular biological applications.
Figures




Similar articles
-
Elimination of cap structures generated by mRNA decay involves the new scavenger mRNA decapping enzyme Aph1/FHIT together with DcpS.Nucleic Acids Res. 2015 Jan;43(1):482-92. doi: 10.1093/nar/gku1251. Epub 2014 Nov 28. Nucleic Acids Res. 2015. PMID: 25432955 Free PMC article.
-
Crystal structures of human DcpS in ligand-free and m7GDP-bound forms suggest a dynamic mechanism for scavenger mRNA decapping.J Mol Biol. 2005 Apr 8;347(4):707-18. doi: 10.1016/j.jmb.2005.01.062. J Mol Biol. 2005. PMID: 15769464
-
Decapping Scavenger (DcpS) enzyme: advances in its structure, activity and roles in the cap-dependent mRNA metabolism.Biochim Biophys Acta. 2014 Jun;1839(6):452-62. doi: 10.1016/j.bbagrm.2014.04.007. Epub 2014 Apr 15. Biochim Biophys Acta. 2014. PMID: 24742626 Review.
-
Molecular basis of the selective processing of short mRNA substrates by the DcpS mRNA decapping enzyme.Proc Natl Acad Sci U S A. 2020 Aug 11;117(32):19237-19244. doi: 10.1073/pnas.2009362117. Epub 2020 Jul 28. Proc Natl Acad Sci U S A. 2020. PMID: 32723815 Free PMC article.
-
Decapping the message: a beginning or an end.Biochem Soc Trans. 2006 Feb;34(Pt 1):35-8. doi: 10.1042/BST20060035. Biochem Soc Trans. 2006. PMID: 16246173 Review.
Cited by
-
mRNA-Decapping Associated DcpS Enzyme Controls Critical Steps of Neuronal Development.Cereb Cortex. 2022 Mar 30;32(7):1494-1507. doi: 10.1093/cercor/bhab302. Cereb Cortex. 2022. PMID: 34467373 Free PMC article.
-
Biochemical characterization of mRNA capping enzyme from Faustovirus.RNA. 2023 Nov;29(11):1803-1817. doi: 10.1261/rna.079738.123. Epub 2023 Aug 25. RNA. 2023. PMID: 37625853 Free PMC article.
-
CompasSeq: epitranscriptome-wide percentage assessment of metabolite-capped RNA at the transcript resolution.Nat Commun. 2025 Jul 20;16(1):6676. doi: 10.1038/s41467-025-61697-y. Nat Commun. 2025. PMID: 40683890 Free PMC article.
-
Prognostic Signature and Tumor Immune Landscape of N7-Methylguanosine-Related lncRNAs in Hepatocellular Carcinoma.Front Genet. 2022 Jul 22;13:906496. doi: 10.3389/fgene.2022.906496. eCollection 2022. Front Genet. 2022. PMID: 35938009 Free PMC article.
-
Fast and Efficient Design of Deep Neural Networks for Predicting N7-Methylguanosine Sites Using autoBioSeqpy.ACS Omega. 2023 May 23;8(22):19728-19740. doi: 10.1021/acsomega.3c01371. eCollection 2023 Jun 6. ACS Omega. 2023. PMID: 37305295 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

