Arabidopsis IPGA1 is a microtubule-associated protein essential for cell expansion during petal morphogenesis
- PMID: 31198941
- PMCID: PMC6793458
- DOI: 10.1093/jxb/erz284
Arabidopsis IPGA1 is a microtubule-associated protein essential for cell expansion during petal morphogenesis
Abstract
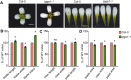
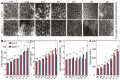
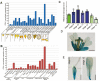
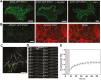
Unlike animal cells, plant cells do not possess centrosomes that serve as microtubule organizing centers; how microtubule arrays are organized throughout plant morphogenesis remains poorly understood. We report here that Arabidopsis INCREASED PETAL GROWTH ANISOTROPY 1 (IPGA1), a previously uncharacterized microtubule-associated protein, regulates petal growth and shape by affecting cortical microtubule organization. Through a genetic screen, we showed that IPGA1 loss-of-function mutants displayed a phenotype of longer and narrower petals, as well as increased anisotropic cell expansion of the petal epidermis in the late phases of flower development. Map-based cloning studies revealed that IPGA1 encodes a previously uncharacterized protein that colocalizes with and directly binds to microtubules. IPGA1 plays a negative role in the organization of cortical microtubules into parallel arrays oriented perpendicular to the axis of cell elongation, with the ipga1-1 mutant displaying increased microtubule ordering in petal abaxial epidermal cells. The IPGA1 family is conserved among land plants and its homologs may have evolved to regulate microtubule organization. Taken together, our findings identify IPGA1 as a novel microtubule-associated protein and provide significant insights into IPGA1-mediated microtubule organization and petal growth anisotropy.
Keywords: Arabidopsis; INCREASED PETAL GROWTH ANISOTROPY 1 (IPGA1); cortical microtubule; growth anisotropy; microtubule-associated protein; petal.
© The Author(s) 2019. Published by Oxford University Press on behalf of the Society for Experimental Biology.
Figures


Similar articles
-
Cortical Microtubule Organization during Petal Morphogenesis in Arabidopsis.Int J Mol Sci. 2019 Oct 3;20(19):4913. doi: 10.3390/ijms20194913. Int J Mol Sci. 2019. PMID: 31623377 Free PMC article. Review.
-
The IPGA1-ANGUSTIFOLIA module regulates microtubule organisation and pavement cell shape in Arabidopsis.New Phytol. 2022 Nov;236(4):1310-1325. doi: 10.1111/nph.18433. Epub 2022 Sep 1. New Phytol. 2022. PMID: 35975703
-
SPIKE1 Activates ROP GTPase to Modulate Petal Growth and Shape.Plant Physiol. 2016 Sep;172(1):358-71. doi: 10.1104/pp.16.00788. Epub 2016 Jul 20. Plant Physiol. 2016. PMID: 27440754 Free PMC article.
-
Spatio-temporal orientation of microtubules controls conical cell shape in Arabidopsis thaliana petals.PLoS Genet. 2017 Jun 23;13(6):e1006851. doi: 10.1371/journal.pgen.1006851. eCollection 2017 Jun. PLoS Genet. 2017. PMID: 28644898 Free PMC article.
-
Building beauty: Understanding how hormone signaling regulates petal patterning and morphogenesis.Plant J. 2025 Mar;121(6):e70101. doi: 10.1111/tpj.70101. Plant J. 2025. PMID: 40106266 Free PMC article. Review.
Cited by
-
Cadmium toxicity: its' uptake and retaliation by plant defence system and ja signaling.Biometals. 2024 Aug;37(4):755-772. doi: 10.1007/s10534-023-00569-8. Epub 2024 Jan 11. Biometals. 2024. PMID: 38206521 Review.
-
Microtubule Regulation in Plants: From Morphological Development to Stress Adaptation.Biomolecules. 2023 Mar 30;13(4):627. doi: 10.3390/biom13040627. Biomolecules. 2023. PMID: 37189374 Free PMC article. Review.
-
The E3 ligase MREL57 modulates microtubule stability and stomatal closure in response to ABA.Nat Commun. 2021 Apr 12;12(1):2181. doi: 10.1038/s41467-021-22455-y. Nat Commun. 2021. PMID: 33846350 Free PMC article.
-
Cortical Microtubule Organization during Petal Morphogenesis in Arabidopsis.Int J Mol Sci. 2019 Oct 3;20(19):4913. doi: 10.3390/ijms20194913. Int J Mol Sci. 2019. PMID: 31623377 Free PMC article. Review.
-
Sculpting the surface: Structural patterning of plant epidermis.iScience. 2021 Oct 25;24(11):103346. doi: 10.1016/j.isci.2021.103346. eCollection 2021 Nov 19. iScience. 2021. PMID: 34820605 Free PMC article. Review.
References
-
- Baskin TI. 2001. On the alignment of cellulose microfibrils by cortical microtubules: a review and a model. Protoplasma 215, 150–171. - PubMed
-
- Baskin TI. 2005. Anisotropic expansion of the plant cell wall. Annual Review of Cell and Developmental Biology 21, 203–222. - PubMed
-
- Baskin TI, Wilson JE, Cork A, Williamson RE. 1994. Morphology and microtubule organization in Arabidopsis roots exposed to oryzalin or taxol. Plant & Cell Physiology 35, 935–942. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

