Glypican 6 is a putative biomarker for metastatic progression of cutaneous melanoma
- PMID: 31199813
- PMCID: PMC6568403
- DOI: 10.1371/journal.pone.0218067
Glypican 6 is a putative biomarker for metastatic progression of cutaneous melanoma
Abstract
Due to the poor prognosis of advanced metastatic melanoma, it is crucial to find early biomarkers that help identify which melanomas will metastasize. By comparing the gene expression data from primary and cutaneous melanoma samples from The Cancer Genome Atlas (TCGA), we identified GPC6 among a set of genes whose expression levels can distinguish between primary melanoma and regional cutaneous/subcutaneous metastases. Glypicans are thought to play a role in tumor growth by regulating the signaling pathways of Wnt, Hedgehogs, fibroblast growth factors (FGFs), and bone morphogenetic proteins (BMPs). We showed that GPC6 expression was up-regulated in a melanoma cell line compared to normal melanocytes and in metastatic melanoma compared to primary melanoma. Furthermore, GPC6 expression was positively correlated with genes largely involved in cell adhesion and migration in both melanoma samples and in RNA-seq samples from other TCGA tumors. Our results suggest that GPC6 may play a role in tumor metastatic progression. In TCGA melanoma samples, we also showed that GPC6 expression was negatively correlated with miR-509-3p, which has previously been shown to function as a tumor suppressor in various cancer cell lines. We overexpressed miR-509-3p in A375 melanoma cells and showed that GPC6 expression was significantly suppressed. This result suggested that GPC6 was a putative target of miR-509-3p in melanoma. Together, our findings identified GPC6 as an early biomarker for melanoma metastatic progression, one that can be regulated by miR-509-3p.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
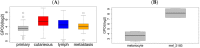



Similar articles
-
Genome-Wide Screen for MicroRNAs Reveals a Role for miR-203 in Melanoma Metastasis.J Invest Dermatol. 2018 Apr;138(4):882-892. doi: 10.1016/j.jid.2017.09.049. Epub 2018 Feb 13. J Invest Dermatol. 2018. PMID: 29104160
-
Regulation of RUNX3 tumor suppressor gene expression in cutaneous melanoma.Clin Cancer Res. 2009 May 1;15(9):2988-94. doi: 10.1158/1078-0432.CCR-08-3172. Epub 2009 Mar 31. Clin Cancer Res. 2009. PMID: 19336521 Free PMC article.
-
Identifying FBLN1 (Gene ID: 2192) as a Potential Melanoma Biomarker for Melanoma based on an Analysis of microRNA Expression Profiles in the GEO and TCGA Databases.Genet Test Mol Biomarkers. 2021 Jan;25(1):68-78. doi: 10.1089/gtmb.2020.0274. Genet Test Mol Biomarkers. 2021. PMID: 33470885 Free PMC article.
-
MicroRNAs in malignant melanoma.Clin Biochem. 2013 Jul;46(10-11):909-17. doi: 10.1016/j.clinbiochem.2013.01.008. Epub 2013 Jan 27. Clin Biochem. 2013. PMID: 23360785 Review.
-
Mutation and expression of TP53 in malignant melanomas.Recent Results Cancer Res. 1995;139:137-54. doi: 10.1007/978-3-642-78771-3_10. Recent Results Cancer Res. 1995. PMID: 7597286 Review.
Cited by
-
Distinct diagnostic and prognostic values of Glypicans gene expression in patients with hepatocellular carcinoma.BMC Cancer. 2021 Apr 26;21(1):462. doi: 10.1186/s12885-021-08104-z. BMC Cancer. 2021. PMID: 33902495 Free PMC article.
-
Non-coding RNAs in skin cancers:Biological roles and molecular mechanisms.Front Pharmacol. 2022 Aug 10;13:934396. doi: 10.3389/fphar.2022.934396. eCollection 2022. Front Pharmacol. 2022. PMID: 36034860 Free PMC article. Review.
-
A Network of Circular RNA and MicroRNA Sequencing Provides Insights into Pigment Deposition of Changshun Blue Eggshell Chickens.Genes (Basel). 2024 Jun 19;15(6):812. doi: 10.3390/genes15060812. Genes (Basel). 2024. PMID: 38927747 Free PMC article.
-
Biological characteristics and clinical management of uveal and conjunctival melanoma.Oncol Res. 2024 Jul 17;32(8):1265-1285. doi: 10.32604/or.2024.048437. eCollection 2024. Oncol Res. 2024. PMID: 39055896 Free PMC article. Review.
-
The Role of Glypicans in Cancer Progression and Therapy.J Histochem Cytochem. 2020 Dec;68(12):841-862. doi: 10.1369/0022155420933709. Epub 2020 Jul 6. J Histochem Cytochem. 2020. PMID: 32623934 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

