Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development
- PMID: 31200566
- PMCID: PMC6627697
- DOI: 10.3390/cells8060583
Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development
Abstract
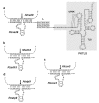
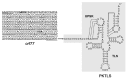
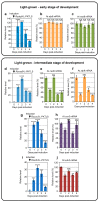
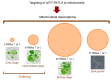
We address here organellar genetic regulation and intercompartment genome coordination. We developed earlier a strategy relying on a tRNA-like shuttle to mediate import of nuclear transgene-encoded custom RNAs into mitochondria in plants. In the present work, we used this strategy to drive trans-cleaving hammerhead ribozymes into the organelles, to knock down specific mitochondrial RNAs and analyze the regulatory impact. In a similar approach, the tRNA mimic was used to import into mitochondria in Arabidopsis thaliana the orf77, an RNA associated with cytoplasmic male sterility in maize and possessing sequence identities with the atp9 mitochondrial RNA. In both cases, inducible expression of the transgenes allowed to characterise early regulation and signaling responses triggered by these respective manipulations of the organellar transcriptome. The results imply that the mitochondrial transcriptome is tightly controlled by a "buffering" mechanism at the early and intermediate stages of plant development, a control that is released at later stages. On the other hand, high throughput analyses showed that knocking down a specific mitochondrial mRNA triggered a retrograde signaling and an anterograde nuclear transcriptome response involving a series of transcription factor genes and small RNAs. Our results strongly support transcriptome coordination mechanisms within the organelles and between the organelles and the nucleus.
Keywords: RNA trafficking; anterograde regulation; cytoplasmic male sterility (CMS); plant mitochondria; retrograde regulation; ribozyme; signaling.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Mitochondrial transport of catalytic RNAs and targeting of the organellar transcriptome in human cells.J Mol Cell Biol. 2024 Jan 17;15(8):mjad051. doi: 10.1093/jmcb/mjad051. J Mol Cell Biol. 2024. PMID: 37591617 Free PMC article.
-
Organelle trafficking of chimeric ribozymes and genetic manipulation of mitochondria.Nucleic Acids Res. 2011 Nov;39(21):9262-74. doi: 10.1093/nar/gkr580. Epub 2011 Jul 17. Nucleic Acids Res. 2011. PMID: 21768127 Free PMC article.
-
A restorer-of-fertility like pentatricopeptide repeat gene directs ribonucleolytic processing within the coding sequence of rps3-rpl16 and orf240a mitochondrial transcripts in Arabidopsis thaliana.Plant J. 2014 Apr;78(1):134-45. doi: 10.1111/tpj.12463. Epub 2014 Mar 12. Plant J. 2014. PMID: 24506331
-
Genome barriers between nuclei and mitochondria exemplified by cytoplasmic male sterility.Plant Cell Physiol. 2008 Oct;49(10):1484-94. doi: 10.1093/pcp/pcn102. Epub 2008 Jul 14. Plant Cell Physiol. 2008. PMID: 18625609 Free PMC article. Review.
-
Mitochondrial retrograde regulation tuning fork in nuclear genes expressions of higher plants.J Genet Genomics. 2008 Feb;35(2):65-71. doi: 10.1016/S1673-8527(08)60010-7. J Genet Genomics. 2008. PMID: 18407052 Review.
Cited by
-
Mitochondrial transport of catalytic RNAs and targeting of the organellar transcriptome in human cells.J Mol Cell Biol. 2024 Jan 17;15(8):mjad051. doi: 10.1093/jmcb/mjad051. J Mol Cell Biol. 2024. PMID: 37591617 Free PMC article.
-
Noncoding RNA: An Insight into Chloroplast and Mitochondrial Gene Expressions.Life (Basel). 2021 Jan 13;11(1):49. doi: 10.3390/life11010049. Life (Basel). 2021. PMID: 33450961 Free PMC article. Review.
-
Triple-localized WHIRLY2 Influences Leaf Senescence and Silique Development via Carbon Allocation.Plant Physiol. 2020 Nov;184(3):1348-1362. doi: 10.1104/pp.20.00832. Epub 2020 Sep 8. Plant Physiol. 2020. PMID: 32900978 Free PMC article.
-
Differentially Expressed Genes Shared by Two Distinct Cytoplasmic Male Sterility (CMS) Types of Silene vulgaris Suggest the Importance of Oxidative Stress in Pollen Abortion.Cells. 2020 Dec 16;9(12):2700. doi: 10.3390/cells9122700. Cells. 2020. PMID: 33339225 Free PMC article.
-
Nitric oxide promotes energy metabolism and protects mitochondrial DNA in peaches during cold storage.Front Plant Sci. 2022 Oct 6;13:970303. doi: 10.3389/fpls.2022.970303. eCollection 2022. Front Plant Sci. 2022. PMID: 36275543 Free PMC article.
References
-
- Holec S., Lange H., Kühn K., Alioua M., Börner T., Gagliardi D. Relaxed transcription in Arabidopsis mitochondria is counterbalanced by RNA stability control mediated by polyadenylation and polynucleotide phosphorylase. Mol. Cell. Biol. 2006;26:2869–2876. doi: 10.1128/MCB.26.7.2869-2876.2006. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

