The cell biology of the hepatocyte: A membrane trafficking machine
- PMID: 31201265
- PMCID: PMC6605791
- DOI: 10.1083/jcb.201903090
The cell biology of the hepatocyte: A membrane trafficking machine
Abstract
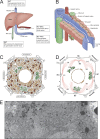
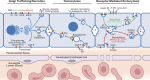
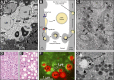
The liver performs numerous vital functions, including the detoxification of blood before access to the brain while simultaneously secreting and internalizing scores of proteins and lipids to maintain appropriate blood chemistry. Furthermore, the liver also synthesizes and secretes bile to enable the digestion of food. These diverse attributes are all performed by hepatocytes, the parenchymal cells of the liver. As predicted, these cells possess a remarkably well-developed and complex membrane trafficking machinery that is dedicated to moving specific cargos to their correct cellular locations. Importantly, while most epithelial cells secrete nascent proteins directionally toward a single lumen, the hepatocyte secretes both proteins and bile concomitantly at its basolateral and apical domains, respectively. In this Beyond the Cell review, we will detail these central features of the hepatocyte and highlight how membrane transport processes play a key role in healthy liver function and how they are affected by disease.
© 2019 Schulze et al.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

