The Properties of 5-Methyltetrahydrofolate Dehydrogenase (MetF1) and Its Role in the Tetrahydrofolate-Dependent Dicamba Demethylation System in Rhizorhabdus dicambivorans Ndbn-20
- PMID: 31209079
- PMCID: PMC6689297
- DOI: 10.1128/JB.00096-19
The Properties of 5-Methyltetrahydrofolate Dehydrogenase (MetF1) and Its Role in the Tetrahydrofolate-Dependent Dicamba Demethylation System in Rhizorhabdus dicambivorans Ndbn-20
Abstract
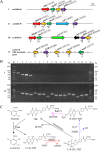
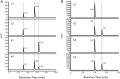
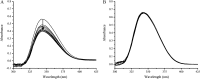
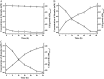
The herbicide dicamba is initially degraded via the tetrahydrofolate (THF)-dependent demethylation system in Rhizorhabdus dicambivorans Ndbn-20. Two THF-dependent dicamba methyltransferase gene clusters, scaffold 50 and scaffold 66, were found in the genome of strain Ndbn-20. Each cluster contains a dicamba methyltransferase gene and three THF metabolism-related genes, namely, metF (coding for 5,10-CH2-THF reductase), folD (coding for 5,10-CH2-THF dehydrogenase-5,10-methenyl-THF cyclohydrolase), and purU (coding for 10-formyl-THF deformylase). In this study, reverse transcription-PCR (RT-PCR) results showed that only genes in scaffold 66, not those in scaffold 50, were transcribed in dicamba-cultured cells. The metF gene of scaffold 66 (metF1) was expressed in Escherichia coli BL21(DE3), and the product was purified as a His6-tagged protein. Purified MetF1 was found to be a monomer and exhibited 5-CH3-THF dehydrogenase activity in vitro The kcat and Km for 5-CH3-THF were 0.23 s-1 and 16.48 μM, respectively. However, 5,10-CH2-THF reductase activity was not detected for MetF1 under the conditions tested. Gene disruption results showed that metF1 is essential for dicamba degradation, whereas folD1 is dispensable.IMPORTANCE There are several THF-dependent methyltransferase genes and THF-metabolic genes in the genome of R. dicambivorans Ndbn-20; however, which genes are involved in dicamba demethylation and the mechanism underlying THF regeneration remain unknown. This study revealed that scaffold 66 is responsible for dicamba demethylation and that MetF1 physiologically catalyzes the dehydrogenation of 5-CH3-THF to 5,10-CH2-THF in the THF-dependent dicamba demethylation system in R. dicambivorans Ndbn-20. Furthermore, the results showed that MetF1 differs from previously characterized MetF in phylogenesis, biochemical properties, and catalytic activity; e.g., MetF1 in vitro did not show 5,10-CH2-THF reductase activity, which is the physiological function of Escherichia coli MetF. This study provides new insights into the mechanism of the THF-dependent methyltransferase system.
Keywords: 5-CH3-THF dehydrogenase activity; MetF1; THF regeneration pathway; THF-dependent dicamba demethylation system; enzymatic characteristics; gene disruption.
Copyright © 2019 American Society for Microbiology.
Figures


References
-
- Yoshikata T, Suzuki K, Kamimura N, Namiki M, Hishiyama S, Araki T, Kasai D, Otsuka Y, Nakamura M, Fukuda M, Katayama Y, Masai E. 2014. Three-component O-demethylase system essential for catabolism of a lignin-derived biphenyl compound in Sphingobium sp. strain SYK-6. Appl Environ Microbiol 80:7142–7153. doi: 10.1128/AEM.02236-14. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

