Conserved properties of genetic architecture of renal and fat transcriptomes in rat models of insulin resistance
- PMID: 31213483
- PMCID: PMC6679378
- DOI: 10.1242/dmm.038539
Conserved properties of genetic architecture of renal and fat transcriptomes in rat models of insulin resistance
Abstract
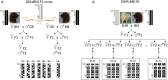
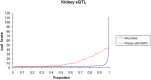
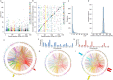
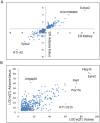
To define renal molecular mechanisms that are affected by permanent hyperglycaemia and might promote phenotypes relevant to diabetic nephropathy, we carried out linkage analysis of genome-wide gene transcription in the kidneys of F2 offspring from the Goto-Kakizaki (GK) rat model of type 2 diabetes and normoglycaemic Brown Norway (BN) rats. We mapped 2526 statistically significant expression quantitative trait loci (eQTLs) in the cross. More than 40% of eQTLs mapped in the close vicinity of the linked transcripts, underlying possible cis-regulatory mechanisms of gene expression. We identified eQTL hotspots on chromosomes 5 and 9 regulating the expression of 80-165 genes, sex or cross direction effects, and enriched metabolic and immunological processes by segregating GK alleles. Comparative analysis with adipose tissue eQTLs in the same cross showed that 496 eQTLs, in addition to the top enriched biological pathways, are conserved in the two tissues. Extensive similarities in eQTLs mapped in the GK rat and in the spontaneously hypertensive rat (SHR) suggest a common aetiology of disease phenotypes common to the two strains, including insulin resistance, which is a prominent pathophysiological feature in both GK rats and SHRs. Our data shed light on shared and tissue-specific molecular mechanisms that might underlie aetiological aspects of insulin resistance in the context of spontaneously occurring hyperglycaemia and hypertension.
Keywords: Diabetes mellitus; Goto-Kakizaki rat; SNP; Spontaneously hypertensive rat; Systems genetics; Transcriptome; eQTL.
© 2019. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures
Similar articles
-
Topological analysis of metabolic networks integrating co-segregating transcriptomes and metabolomes in type 2 diabetic rat congenic series.Genome Med. 2016 Sep 30;8(1):101. doi: 10.1186/s13073-016-0352-6. Genome Med. 2016. PMID: 27716393 Free PMC article.
-
Enhanced insulin secretion and cholesterol metabolism in congenic strains of the spontaneously diabetic (Type 2) Goto Kakizaki rat are controlled by independent genetic loci in rat chromosome 8.Diabetologia. 2004 Jun;47(6):1096-106. doi: 10.1007/s00125-004-1416-5. Epub 2004 May 26. Diabetologia. 2004. PMID: 15164172
-
Genetic control of plasma lipid levels in a cross derived from normoglycaemic Brown Norway and spontaneously diabetic Goto-Kakizaki rats.Diabetologia. 2006 Nov;49(11):2679-88. doi: 10.1007/s00125-006-0396-z. Epub 2006 Sep 16. Diabetologia. 2006. PMID: 16983556
-
Genomic regulation of type 2 diabetes endophenotypes: Contribution from genetic studies in the Goto-Kakizaki rat.Biochimie. 2017 Dec;143:56-65. doi: 10.1016/j.biochi.2017.08.012. Epub 2017 Aug 24. Biochimie. 2017. PMID: 28843612 Review.
-
Mapping genetic determinants of kidney damage in rat models.Hypertens Res. 2012 Jul;35(7):675-94. doi: 10.1038/hr.2012.77. Epub 2012 May 31. Hypertens Res. 2012. PMID: 22648060 Review.
Cited by
-
What lies beyond 100 years of insulin.Dis Model Mech. 2021 Nov 1;14(11):dmm049361. doi: 10.1242/dmm.049361. Epub 2021 Nov 9. Dis Model Mech. 2021. PMID: 34752619 Free PMC article.
-
lncRNA-mRNA competing endogenous RNA network in IR-hepG2 cells ameliorated by APBBR decreasing ROS levels: a systematic analysis.PeerJ. 2020 Feb 24;8:e8604. doi: 10.7717/peerj.8604. eCollection 2020. PeerJ. 2020. PMID: 32140303 Free PMC article.
-
The regulatory landscape of multiple brain regions in outbred heterogeneous stock rats.Nucleic Acids Res. 2022 Oct 28;50(19):10882-10895. doi: 10.1093/nar/gkac912. Nucleic Acids Res. 2022. PMID: 36263809 Free PMC article.
-
Dominant gut Prevotella copri in gastrectomised non-obese diabetic Goto-Kakizaki rats improves glucose homeostasis through enhanced FXR signalling.Diabetologia. 2020 Jun;63(6):1223-1235. doi: 10.1007/s00125-020-05122-7. Epub 2020 Mar 16. Diabetologia. 2020. PMID: 32173762 Free PMC article.
References
-
- Aitman T. J., Glazier A. M., Wallace C. A., Cooper L. D., Norsworthy P. J., Wahid F. N., Al-Majali K. M., Trembling P. M., Mann C. J., Shoulders C. C. et al. (1999). Identification of Cd36 (Fat) as an insulin-resistance gene causing defective fatty acid and glucose metabolism in hypertensive rats. Nat. Genet. 21, 76-83. 10.1038/5013 - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

