Screening methods for detection of ancient Mycobacterium tuberculosis complex fingerprints in next-generation sequencing data derived from skeletal samples
- PMID: 31220249
- PMCID: PMC6586198
- DOI: 10.1093/gigascience/giz065
Screening methods for detection of ancient Mycobacterium tuberculosis complex fingerprints in next-generation sequencing data derived from skeletal samples
Abstract
Background: Recent advances in ancient DNA studies, especially in increasing isolated DNA yields and quality, have opened the possibility of analysis of ancient host microbiome. However, such pitfalls as spurious identification of pathogens based on fragmentary data or environmental contamination could lead to incorrect epidaemiological conclusions. Within the Mycobacterium genus, Mycobacterium tuberculosis complex members responsible for tuberculosis share up to ∼99% genomic sequence identity, while other more distantly related Mycobacteria other than M. tuberculosis can be causative agents for pulmonary diseases or soil dwellers. Therefore, reliable determination of species complex is crucial for interpretation of sequencing results.
Results: Here we present a novel bioinformatical approach, used for screening of ancient tuberculosis in sequencing data, derived from 28 individuals (dated 4400-4000 and 3100-2900 BC) from central Poland. We demonstrate that cost-effective next-generation screening sequencing data (∼20M reads per sample) could yield enough information to provide statistically supported identification of probable ancient disease cases.
Conclusions: Application of appropriate bioinformatic tools, including an unbiased selection of genomic alignment targets for species specificity, makes it possible to extract valid data from full-sample sequencing results (without subjective targeted enrichment procedures). This approach broadens the potential scope of palaeoepidaemiology both to older, suboptimally preserved samples and to pathogens with difficult intrageneric taxonomy.
Keywords: NGS; aTB; ancient DNA; ancient tuberculosis.
© The Author(s) 2019. Published by Oxford University Press.
Figures


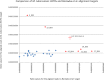
References
-
- Barrett R, Kuzawa CW, McDade T, et al.. Emerging and re-emerging infectious diseases: the third epidemiologic transition. Annu Rev Anthropol. 1998;27(1):247–71.
-
- Armelagos GJ, Cohen MN. Paleopathology at the Origins of Agriculture. Orlando, FL: Academic Press; 1984.
-
- Armelagos GJ, Goodman AH, Jacobs KH. The origins of agriculture: population growth during a period of declining health. Popul Environ. 1991;13(1):9–22.
-
- Ortner DJ, Putschar W. Identification of Pathological Conditions in Human Skeletal Remains. Smithsonian; 1985.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

