Trapping a somatic endogenous retrovirus into a germline piRNA cluster immunizes the germline against further invasion
- PMID: 31227013
- PMCID: PMC6587276
- DOI: 10.1186/s13059-019-1736-x
Trapping a somatic endogenous retrovirus into a germline piRNA cluster immunizes the germline against further invasion
Abstract
Background: For species survival, the germline must faithfully transmit genetic information to the progeny. Transposable elements (TEs) constitute a significant threat to genome stability due to their mobility. In the metazoan germline, their mobilization is limited by a class of small RNAs called PIWI-interacting RNAs (piRNAs) produced by dedicated genomic loci called piRNA clusters. Although the piRNA pathway is an adaptive genomic immunity system, it remains unclear how the germline gains protection from a new transposon invasion.
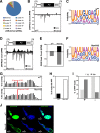
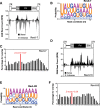
Results: To address this question, we analyze Drosophila melanogaster lines harboring a deletion within flamenco, a major piRNA cluster specifically expressed in somatic follicular cells. This deletion leads to derepression of the retrotransposon ZAM in the somatic follicular cells and subsequent germline genome invasion. In this mutant line, we identify de novo production of sense and antisense ZAM-derived piRNAs that display a germinal molecular signature. These piRNAs originated from a new ZAM insertion into a germline dual-strand piRNA cluster and silence ZAM expression specifically in germ cells. Finally, we find that ZAM trapping in a germinal piRNA cluster is a frequent event that occurs early during the isolation of the mutant line.
Conclusions: Transposons can hijack the host developmental process to propagate whenever their silencing is lost. Here, we show that the germline can protect itself by trapping invading somatic-specific TEs into germline piRNA clusters. This is the first demonstration of "auto-immunization" of a germline endangered by mobilization of a surrounding somatic TE.
Keywords: Drosophila; Genome stability; Germline; Inheritance; Transposable elements; piRNA cluster; piRNAs.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
The Tudor domain protein Tapas, a homolog of the vertebrate Tdrd7, functions in the piRNA pathway to regulate retrotransposons in germline of Drosophila melanogaster.BMC Biol. 2014 Oct 6;12:61. doi: 10.1186/s12915-014-0061-9. BMC Biol. 2014. PMID: 25287931 Free PMC article.
-
piRNA and Transposon Dynamics in Drosophila: A Female Story.Genome Biol Evol. 2020 Jun 1;12(6):931-947. doi: 10.1093/gbe/evaa094. Genome Biol Evol. 2020. PMID: 32396626 Free PMC article.
-
Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary.Cell. 2009 May 1;137(3):522-35. doi: 10.1016/j.cell.2009.03.040. Epub 2009 Apr 23. Cell. 2009. PMID: 19395010 Free PMC article.
-
piRNA Defense Against Endogenous Retroviruses.Viruses. 2024 Nov 9;16(11):1756. doi: 10.3390/v16111756. Viruses. 2024. PMID: 39599869 Free PMC article. Review.
-
The piRNA pathway in Drosophila ovarian germ and somatic cells.Proc Jpn Acad Ser B Phys Biol Sci. 2020;96(1):32-42. doi: 10.2183/pjab.96.003. Proc Jpn Acad Ser B Phys Biol Sci. 2020. PMID: 31932527 Free PMC article. Review.
Cited by
-
Rapid evolutionary diversification of the flamenco locus across simulans clade Drosophila species.PLoS Genet. 2023 Aug 29;19(8):e1010914. doi: 10.1371/journal.pgen.1010914. eCollection 2023 Aug. PLoS Genet. 2023. PMID: 37643184 Free PMC article.
-
Small RNA Deep Sequencing Identifies a Unique miRNA Signature Released in Serum Exosomes in a Mouse Model of Sjögren's Syndrome.Front Immunol. 2020 Jul 17;11:1475. doi: 10.3389/fimmu.2020.01475. eCollection 2020. Front Immunol. 2020. PMID: 32849505 Free PMC article.
-
What Are the Functional Roles of Piwi Proteins and piRNAs in Insects?Insects. 2023 Feb 14;14(2):187. doi: 10.3390/insects14020187. Insects. 2023. PMID: 36835756 Free PMC article. Review.
-
Retrotransposons Down- and Up-Regulation in Aging Somatic Tissues.Cells. 2021 Dec 28;11(1):79. doi: 10.3390/cells11010079. Cells. 2021. PMID: 35011640 Free PMC article.
-
Reactivation of a somatic errantivirus and germline invasion in Drosophila ovaries.Nat Commun. 2023 Sep 29;14(1):6096. doi: 10.1038/s41467-023-41733-5. Nat Commun. 2023. PMID: 37773253 Free PMC article.
References
-
- Aravin A, Gaidatzis D, Pfeffer S, Lagos-Quintana M, Landgraf P, Iovino N, et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nat Cell Biol. 2006;442(7099):203–207. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

