In Silico Insights into HIV-1 Vpu-Tetherin Interactions and Its Mutational Counterparts
- PMID: 31234536
- PMCID: PMC6631454
- DOI: 10.3390/medsci7060074
In Silico Insights into HIV-1 Vpu-Tetherin Interactions and Its Mutational Counterparts
Abstract
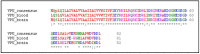
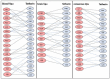
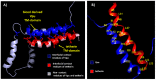
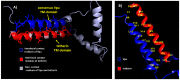
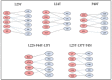
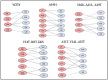
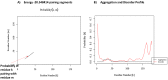
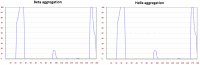
Tetherin, an interferon-induced host protein encoded by the bone marrow stromal antigen 2 (BST2/CD317/HM1.24) gene, is involved in obstructing the release of many retroviruses and other enveloped viruses by cross-linking the budding virus particles to the cell surface. This activity is antagonized in the case of human immunodeficiency virus (HIV)-1 wherein its accessory protein Viral Protein U (Vpu) interacts with tetherin, causing its downregulation from the cell surface. Vpu and tetherin connect through their transmembrane (TM) domains, culminating into events leading to tetherin degradation by recruitment of β-TrCP2. However, mutations in the TM domains of both proteins are reported to act as a resistance mechanism to Vpu countermeasure impacting tetherin's sensitivity towards Vpu but retaining its antiviral activity. Our study illustrates the binding aspects of blood-derived, brain-derived, and consensus HIV-1 Vpu with tetherin through protein-protein docking. The analysis of the bound complexes confirms the blood-derived Vpu-tetherin complex to have the best binding affinity as compared to other two. The mutations in tetherin and Vpu are devised computationally and are subjected to protein-protein interactions. The complexes are tested for their binding affinities, residue connections, hydrophobic forces, and, finally, the effect of mutation on their interactions. The single point mutations in tetherin at positions L23Y, L24T, and P40T, and triple mutations at {L22S, F44Y, L37I} and {L23T, L37T, T45I}, while single point mutations in Vpu at positions A19H and W23Y and triplet of mutations at {V10K, A11L, A19T}, {V14T, I18T, I26S}, and {A11T, V14L, A15T} have revealed no polar contacts with minimal hydrophobic interactions between Vpu and tetherin, resulting in reduced binding affinity. Additionally, we have explored the aggregation potential of tetherin and its association with the brain-derived Vpu protein. This work is a possible step toward an understanding of Vpu-tetherin interactions.
Keywords: HIV-1 Vpu; aggregation potential; tetherin; transmembrane interactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
HIV-1 Vpu Antagonizes CD317/Tetherin by Adaptor Protein-1-Mediated Exclusion from Virus Assembly Sites.J Virol. 2016 Jul 11;90(15):6709-6723. doi: 10.1128/JVI.00504-16. Print 2016 Aug 1. J Virol. 2016. PMID: 27170757 Free PMC article.
-
Tetherin inhibits retrovirus release and is antagonized by HIV-1 Vpu.Nature. 2008 Jan 24;451(7177):425-30. doi: 10.1038/nature06553. Epub 2008 Jan 16. Nature. 2008. PMID: 18200009
-
Vpu-Mediated Counteraction of Tetherin Is a Major Determinant of HIV-1 Interferon Resistance.mBio. 2016 Aug 16;7(4):e00934-16. doi: 10.1128/mBio.00934-16. mBio. 2016. PMID: 27531907 Free PMC article.
-
Transmembrane interactions of HIV-1 Vpu and tetherin.Curr HIV Res. 2012 Jun;10(4):292-7. doi: 10.2174/157016212800792450. Curr HIV Res. 2012. PMID: 22524177 Review.
-
Role of the endosomal ESCRT machinery in HIV-1 Vpu-induced down-regulation of BST2/tetherin.Curr HIV Res. 2012 Jun;10(4):315-20. doi: 10.2174/157016212800792414. Curr HIV Res. 2012. PMID: 22524180 Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources

