Protection of abasic sites during DNA replication by a stable thiazolidine protein-DNA cross-link
- PMID: 31235915
- PMCID: PMC6628887
- DOI: 10.1038/s41594-019-0255-5
Protection of abasic sites during DNA replication by a stable thiazolidine protein-DNA cross-link
Abstract
Abasic (AP) sites are one of the most common DNA lesions that block replicative polymerases. 5-hydroxymethylcytosine binding, embryonic stem cell-specific protein (HMCES) recognizes and processes these lesions in the context of single-stranded DNA (ssDNA). A HMCES DNA-protein cross-link (DPC) intermediate is thought to shield the AP site from endonucleases and error-prone polymerases. The highly evolutionarily conserved SOS-response associated peptidase (SRAP) domain of HMCES and its Escherichia coli ortholog YedK mediate lesion recognition. Here we uncover the basis of AP site protection by SRAP domains from a crystal structure of the YedK DPC. YedK forms a stable thiazolidine linkage between a ring-opened AP site and the α-amino and sulfhydryl substituents of its amino-terminal cysteine residue. The thiazolidine linkage explains the remarkable stability of the HMCES DPC, its resistance to strand cleavage and the proteolysis requirement for resolution. Furthermore, its structure reveals that HMCES has specificity for AP sites in ssDNA at junctions found when replicative polymerases encounter the AP lesion.
Conflict of interest statement
Competing Interests Statement
The authors declare no conflicts of interest.
Figures

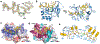
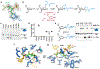
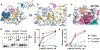
References
-
- Hitomi K, Iwai S & Tainer JA The intricate structural chemistry of base excision repair machinery: implications for DNA damage recognition, removal, and repair. DNA Repair (Amst) 6, 410–28 (2007). - PubMed
Methods-only References
-
- Krokan HE, Saetrom P, Aas PA, Pettersen HS, Kavli B & Slupphaug G Error-free versus mutagenic processing of genomic uracil--relevance to cancer. DNA Repair (Amst) 19, 38–47 (2014). - PubMed
-
- Otwinowski Z & Minor W Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997). - PubMed
-
- Emsley P & Cowtan K Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

