Expanding the Vocabulary of Peptide Signals in Streptococcus mutans
- PMID: 31245303
- PMCID: PMC6563777
- DOI: 10.3389/fcimb.2019.00194
Expanding the Vocabulary of Peptide Signals in Streptococcus mutans
Abstract
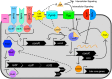
Streptococci, including the dental pathogen Streptococcus mutans, undergo cell-to-cell signaling that is mediated by small peptides to control critical physiological functions such as adaptation to the environment, control of subpopulation behaviors and regulation of virulence factors. One such model pathway is the regulation of genetic competence, controlled by the ComRS signaling system and the peptide XIP. However, recent research in the characterization of this pathway has uncovered novel operons and peptides that are intertwined into its regulation. These discoveries, such as cell lysis playing a critical role in XIP release and importance of bacterial self-sensing during the signaling process, have caused us to reevaluate previous paradigms and shift our views on the true purpose of these signaling systems. The finding of new peptides such as the ComRS inhibitor XrpA and the peptides of the RcrRPQ operon also suggests there may be more peptides hidden in the genomes of streptococci that could play critical roles in the physiology of these organisms. In this review, we summarize the recent findings in S. mutans regarding the integration of other circuits into the ComRS signaling pathway, the true mode of XIP export, and how the RcrRPQ operon controls competence activation. We also look at how new technologies can be used to re-annotate the genome to find new open reading frames that encode peptide signals. Together, this summary of research will allow us to reconsider how we perceive these systems to behave and lead us to expand our vocabulary of peptide signals within the genus Streptococcus.
Keywords: LC-MS/MS; RNA-Seq; Ribo-Seq; bacterial communication; cell-to-cell signaling; genetic competence; peptides; transformation.
Figures

Similar articles
-
Intracellular Signaling by the comRS System in Streptococcus mutans Genetic Competence.mSphere. 2018 Oct 31;3(5):e00444-18. doi: 10.1128/mSphere.00444-18. mSphere. 2018. PMID: 30381353 Free PMC article.
-
Peptides encoded in the Streptococcus mutans RcrRPQ operon are essential for thermotolerance.Microbiology (Reading). 2020 Mar;166(3):306-317. doi: 10.1099/mic.0.000887. Microbiology (Reading). 2020. PMID: 31935187 Free PMC article.
-
Bidirectional signaling in the competence regulatory pathway of Streptococcus mutans.FEMS Microbiol Lett. 2015 Oct;362(19):fnv159. doi: 10.1093/femsle/fnv159. Epub 2015 Sep 10. FEMS Microbiol Lett. 2015. PMID: 26363019 Free PMC article.
-
Quorum sensing and biofilm formation by Streptococcus mutans.Adv Exp Med Biol. 2008;631:178-88. doi: 10.1007/978-0-387-78885-2_12. Adv Exp Med Biol. 2008. PMID: 18792689 Review.
-
The VicRK Two-Component System Regulates Streptococcus mutans Virulence.Curr Issues Mol Biol. 2019;32:167-200. doi: 10.21775/cimb.032.167. Epub 2019 Jun 5. Curr Issues Mol Biol. 2019. PMID: 31166172 Review.
Cited by
-
Small RNA SmsR1 modulates acidogenicity and cariogenic virulence by affecting protein acetylation in Streptococcus mutans.PLoS Pathog. 2024 Apr 15;20(4):e1012147. doi: 10.1371/journal.ppat.1012147. eCollection 2024 Apr. PLoS Pathog. 2024. PMID: 38620039 Free PMC article.
-
Drugs for the Quorum Sensing Inhibition of Oral Biofilm: New Frontiers and Insights in the Treatment of Periodontitis.Pharmaceutics. 2022 Dec 7;14(12):2740. doi: 10.3390/pharmaceutics14122740. Pharmaceutics. 2022. PMID: 36559234 Free PMC article. Review.
-
Direct interactions with commensal streptococci modify intercellular communication behaviors of Streptococcus mutans.ISME J. 2021 Feb;15(2):473-488. doi: 10.1038/s41396-020-00789-7. Epub 2020 Sep 30. ISME J. 2021. PMID: 32999420 Free PMC article.
-
Optimizing the Bioprocesses of Bacteriocin Production in Lacticaseibacillus paracasei HD1.7 by the "Acetate Switch": Novel Insights into the Labor Division Between Energy Metabolism, Quorum Sensing, and Acetate.Foods. 2025 Jul 30;14(15):2691. doi: 10.3390/foods14152691. Foods. 2025. PMID: 40807629 Free PMC article.
-
Structure Activity Relationship Study of the XIP Quorum Sensing Pheromone in Streptococcus mutans Reveal Inhibitors of the Competence Regulon.ACS Chem Biol. 2020 Oct 16;15(10):2833-2841. doi: 10.1021/acschembio.0c00650. Epub 2020 Oct 2. ACS Chem Biol. 2020. PMID: 32946208 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

