Histone Code and Higher-Order Chromatin Folding: A Hypothesis
- PMID: 31245531
- PMCID: PMC6594697
- DOI: 10.18547/gcb.2017.vol3.iss2.e41
Histone Code and Higher-Order Chromatin Folding: A Hypothesis
Abstract
Histone modifications alone or in combination are thought to modulate chromatin structure and function; a concept termed histone code. By combining evidence from several studies, we investigated if the histone code can play a role in higher-order folding of chromatin. Firstly using genomic data, we analyzed associations between histone modifications at the nucleosome level. We could dissect the composition of individual nucleosomes into five predicted clusters of histone modifications. Secondly, by assembling the raw reads of histone modifications at various length scales, we noticed that the histone mark relationships that exist at nucleosome level tend to be maintained at the higher orders of chromatin folding. Recently, a high-resolution imaging study showed that histone marks belonging to three of the five predicted clusters show structurally distinct and anti-correlated chromatin domains at the level of chromosomes. This made us think that the histone code can have a significant impact in the overall compaction of DNA: at the level of nucleosomes, at the level of genes, and finally at the level of chromosomes. As a result, in this article, we put forward a theory where the histone code drives not only the functionality but also the higher-order folding and compaction of chromatin.
Keywords: chromatin folding; chromatin organization; epigenetic regulation; histone code; histone modification; meiosis; nucleosome; super-resolution microscopy.
Conflict of interest statement
CONFLICT OF INTEREST The authors declare no conflict of interest.
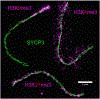
Figures

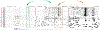

Similar articles
-
Transcriptionally active chromatin loops contain both 'active' and 'inactive' histone modifications that exhibit exclusivity at the level of nucleosome clusters.Epigenetics Chromatin. 2024 Mar 25;17(1):8. doi: 10.1186/s13072-024-00535-9. Epigenetics Chromatin. 2024. PMID: 38528624 Free PMC article.
-
Evidence for the implication of the histone code in building the genome structure.Biosystems. 2018 Feb;164:49-59. doi: 10.1016/j.biosystems.2017.11.005. Epub 2017 Nov 20. Biosystems. 2018. PMID: 29158132 Review.
-
The nucleosome: a little variation goes a long way.Biochem Cell Biol. 2006 Aug;84(4):505-17. doi: 10.1139/o06-085. Biochem Cell Biol. 2006. PMID: 16936823 Review.
-
Hierarchical looping of zigzag nucleosome chains in metaphase chromosomes.Proc Natl Acad Sci U S A. 2016 Feb 2;113(5):1238-43. doi: 10.1073/pnas.1518280113. Epub 2016 Jan 19. Proc Natl Acad Sci U S A. 2016. PMID: 26787893 Free PMC article.
-
Local compartment changes and regulatory landscape alterations in histone H1-depleted cells.Genome Biol. 2015 Dec 23;16:289. doi: 10.1186/s13059-015-0857-0. Genome Biol. 2015. PMID: 26700097 Free PMC article.
Cited by
-
Emerging Contributions of Solid-State NMR Spectroscopy to Chromatin Structural Biology.Front Mol Biosci. 2021 Oct 11;8:741581. doi: 10.3389/fmolb.2021.741581. eCollection 2021. Front Mol Biosci. 2021. PMID: 34708075 Free PMC article. Review.
-
Epigenetic regulation of nuclear processes in fungal plant pathogens.PLoS Pathog. 2023 Aug 3;19(8):e1011525. doi: 10.1371/journal.ppat.1011525. eCollection 2023 Aug. PLoS Pathog. 2023. PMID: 37535497 Free PMC article. Review.
-
Nuclear lamin A/C phosphorylation by loss of androgen receptor leads to cancer-associated fibroblast activation.Nat Commun. 2024 Sep 12;15(1):7984. doi: 10.1038/s41467-024-52344-z. Nat Commun. 2024. PMID: 39266569 Free PMC article.
-
Proteomic variations of esophageal squamous cell carcinoma revealed by combining RNA-seq proteogenomics and G-PTM search strategy.Heliyon. 2020 Aug 29;6(8):e04813. doi: 10.1016/j.heliyon.2020.e04813. eCollection 2020 Aug. Heliyon. 2020. PMID: 32913912 Free PMC article.
-
CTCF-mediated chromatin looping in EGR2 regulation and SUZ12 recruitment critical for peripheral myelination and repair.Nat Commun. 2020 Aug 17;11(1):4133. doi: 10.1038/s41467-020-17955-2. Nat Commun. 2020. PMID: 32807777 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
