Communication codes in developmental signaling pathways
- PMID: 31249008
- PMCID: PMC6602343
- DOI: 10.1242/dev.170977
Communication codes in developmental signaling pathways
Abstract
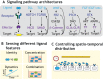
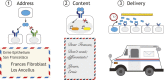
A handful of core intercellular signaling pathways play pivotal roles in a broad variety of developmental processes. It has remained puzzling how so few pathways can provide the precision and specificity of cell-cell communication required for multicellular development. Solving this requires us to quantitatively understand how developmentally relevant signaling information is actively sensed, transformed and spatially distributed by signaling pathways. Recently, single cell analysis and cell-based reconstitution, among other approaches, have begun to reveal the 'communication codes' through which information is represented in the identities, concentrations, combinations and dynamics of extracellular ligands. They have also revealed how signaling pathways decipher these features and control the spatial distribution of signaling in multicellular contexts. Here, we review recent work reporting the discovery and analysis of communication codes and discuss their implications for diverse developmental processes.
Keywords: Communication codes; Pathway architecture; Signal processing.
© 2019. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures





References
-
- Adler M. and Alon U. (2018). Fold-change detection in biological systems. Curr. Opin. Syst. Biol. 8, 81-89. 10.1016/j.coisb.2017.12.005 - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

