A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys
- PMID: 31249312
- PMCID: PMC6597610
- DOI: 10.1038/s41467-019-10861-2
A single-nucleus RNA-sequencing pipeline to decipher the molecular anatomy and pathophysiology of human kidneys
Abstract
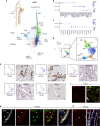
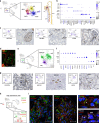
Defining cellular and molecular identities within the kidney is necessary to understand its organization and function in health and disease. Here we demonstrate a reproducible method with minimal artifacts for single-nucleus Droplet-based RNA sequencing (snDrop-Seq) that we use to resolve thirty distinct cell populations in human adult kidney. We define molecular transition states along more than ten nephron segments spanning two major kidney regions. We further delineate cell type-specific expression of genes associated with chronic kidney disease, diabetes and hypertension, providing insight into possible targeted therapies. This includes expression of a hypertension-associated mechano-sensory ion channel in mesangial cells, and identification of proximal tubule cell populations defined by pathogenic expression signatures. Our fully optimized, quality-controlled transcriptomic profiling pipeline constitutes a tool for the generation of healthy and diseased molecular atlases applicable to clinical samples.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Grants and funding
- R01 DK085231/DK/NIDDK NIH HHS/United States
- DK114933/U.S. Department of Health & Human Services | NIH | National Institute of Diabetes and Digestive and Kidney Diseases (National Institute of Diabetes & Digestive & Kidney Diseases)/International
- R01 DK082531/DK/NIDDK NIH HHS/United States
- UG3 DK114933/DK/NIDDK NIH HHS/United States
- U2C DK114886/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

