Brain Molecular Connectivity in Neurodegenerative Diseases: Recent Advances and New Perspectives Using Positron Emission Tomography
- PMID: 31258466
- PMCID: PMC6587303
- DOI: 10.3389/fnins.2019.00617
Brain Molecular Connectivity in Neurodegenerative Diseases: Recent Advances and New Perspectives Using Positron Emission Tomography
Abstract
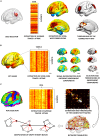
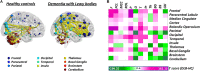
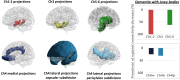
Positron emission tomography (PET) represents a unique molecular tool to get in vivo access to a wide spectrum of biological and neuropathological processes, of crucial relevance for neurodegenerative conditions. Although most PET findings are based on massive univariate approaches, in the last decade the increasing interest in multivariate methods has paved the way to the assessment of unexplored cerebral features, spanning from resting state brain networks to whole-brain connectome properties. Currently, the combination of molecular neuroimaging techniques with multivariate connectivity methods represents one of the most powerful, yet still emerging, approach to achieve novel insights into the pathophysiology of neurodegenerative diseases. In this review, we will summarize the available evidence in the field of PET molecular connectivity, with the aim to provide an overview of how these studies may increase the understanding of the pathogenesis of neurodegenerative diseases, over and above "traditional" structural/functional connectivity studies. Considering the available evidence, a major focus will be represented by molecular connectivity studies using [18F]FDG-PET, today applied in the major neuropathological spectra, from amyloidopathies and tauopathies to synucleinopathies and beyond. Pioneering studies using PET tracers targeting brain neuropathology and neurotransmission systems for connectivity studies will be discussed, their strengths and limitations highlighted with reference to both applied methodology and results interpretation. The most common methods for molecular connectivity assessment will be reviewed, with particular emphasis on the available strategies to investigate molecular connectivity at the single-subject level, of potential relevance for not only research but also diagnostic purposes. Finally, we will highlight possible future perspectives in the field, with reference in particular to newly available PET tracers, which will expand the application of molecular connectivity to new, exciting, unforeseen possibilities.
Keywords: FDG–PET; amyloid PET; brain networks; connectivity; multivariate analysis; neurodegenerative diseases; neurotransmission; tau PET.
Figures
Similar articles
-
Brain Molecular Connectivity in Neurodegenerative Conditions.Brain Sci. 2021 Mar 28;11(4):433. doi: 10.3390/brainsci11040433. Brain Sci. 2021. PMID: 33800680 Free PMC article. Review.
-
A new perspective for advanced positron emission tomography-based molecular imaging in neurodegenerative proteinopathies.Alzheimers Dement. 2019 Aug;15(8):1081-1103. doi: 10.1016/j.jalz.2019.02.004. Epub 2019 Jun 21. Alzheimers Dement. 2019. PMID: 31230910
-
Hybrid PET/MR Imaging and Brain Connectivity.Front Neurosci. 2016 Mar 1;10:64. doi: 10.3389/fnins.2016.00064. eCollection 2016. Front Neurosci. 2016. PMID: 26973446 Free PMC article. Review.
-
Hybrid PET-MRI Applications in Movement Disorders.Int Rev Neurobiol. 2019;144:211-257. doi: 10.1016/bs.irn.2018.10.003. Epub 2018 Nov 8. Int Rev Neurobiol. 2019. PMID: 30638455 Review.
-
Metabolic connectivity: methods and applications.Curr Opin Neurol. 2017 Dec;30(6):677-685. doi: 10.1097/WCO.0000000000000494. Curr Opin Neurol. 2017. PMID: 28914733 Review.
Cited by
-
Comparative test-retest variability of outcome parameters derived from brain [18F]FDG PET studies in non-human primates.PLoS One. 2020 Oct 5;15(10):e0240228. doi: 10.1371/journal.pone.0240228. eCollection 2020. PLoS One. 2020. PMID: 33017429 Free PMC article.
-
Vulnerability of multiple large-scale brain networks in dementia with Lewy bodies.Hum Brain Mapp. 2019 Oct 15;40(15):4537-4550. doi: 10.1002/hbm.24719. Epub 2019 Jul 19. Hum Brain Mapp. 2019. PMID: 31322307 Free PMC article.
-
Functional Brain Networks to Evaluate Treatment Responses in Parkinson's Disease.Neurotherapeutics. 2023 Oct;20(6):1653-1668. doi: 10.1007/s13311-023-01433-w. Epub 2023 Sep 8. Neurotherapeutics. 2023. PMID: 37684533 Free PMC article.
-
In vivo human molecular neuroimaging of dopaminergic vulnerability along the Alzheimer's disease phases.Alzheimers Res Ther. 2021 Nov 12;13(1):187. doi: 10.1186/s13195-021-00925-1. Alzheimers Res Ther. 2021. PMID: 34772450 Free PMC article.
-
Neuromark PET: A multivariate method for estimating whole brain fMRI guided intrinsic networks and connectomes from fMRI and PET data.bioRxiv [Preprint]. 2025 Jan 15:2024.01.10.575131. doi: 10.1101/2024.01.10.575131. bioRxiv. 2025. PMID: 38260682 Free PMC article. Preprint.
References
-
- Ahmed Z., Cooper J., Murray T. K., Garn K., McNaughton S. A., Clarke H., et al. (2014). A novel in vivo model of tau propagation with rapid and progressive neurofibrillary tangle pathology: the pattern of spread is determined by connectivity, not proximity. Acta Neuropathol. 127 667–683. 10.1007/s00401-014-1254-6 - DOI - PMC - PubMed
-
- Albert M. S., DeKosky S. T., Dickson D., Dubois B., Feldman H. H., Fox N. C., et al. (2011). The diagnosis of mild cognitive impairment due to Alzheimer’s disease: recommendations from the National Institute on Aging and Alzheimer’s Association workgroup. Alzheimers Dement. 7 270–279. 10.1016/j.jalz.2011.03.008 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources

