Solvent-dependent segmental dynamics in intrinsically disordered proteins
- PMID: 31259246
- PMCID: PMC6598773
- DOI: 10.1126/sciadv.aax2348
Solvent-dependent segmental dynamics in intrinsically disordered proteins
Abstract
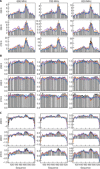
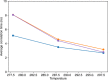
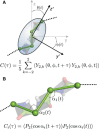
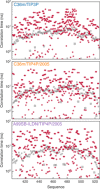
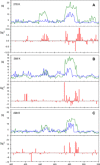
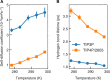
Protein and water dynamics have a synergistic relationship, which is particularly important for intrinsically disordered proteins (IDPs), although the details of this coupling remain poorly understood. Here, we combine temperature-dependent molecular dynamics simulations using different water models with extensive nuclear magnetic resonance (NMR) relaxation to examine the importance of distinct modes of solvent and solute motion for the accurate reproduction of site-specific dynamics in IDPs. We find that water dynamics play a key role in motional processes internal to "segments" of IDPs, stretches of primary sequence that share dynamic properties and behave as discrete dynamic units. We identify a relationship between the time scales of intrasegment dynamics and the lifetime of hydrogen bonds in bulk water. Correct description of these motions is essential for accurate reproduction of protein relaxation. Our findings open important perspectives for understanding the role of hydration water on the behavior and function of IDPs in solution.
Figures
Similar articles
-
Analytical Description of NMR Relaxation Highlights Correlated Dynamics in Intrinsically Disordered Proteins.Angew Chem Int Ed Engl. 2017 Nov 6;56(45):14020-14024. doi: 10.1002/anie.201706740. Epub 2017 Sep 14. Angew Chem Int Ed Engl. 2017. PMID: 28834051
-
Identification of Dynamic Modes in an Intrinsically Disordered Protein Using Temperature-Dependent NMR Relaxation.J Am Chem Soc. 2016 May 18;138(19):6240-51. doi: 10.1021/jacs.6b02424. Epub 2016 May 6. J Am Chem Soc. 2016. PMID: 27112095
-
Development of Charge-Augmented Three-Point Water Model (CAIPi3P) for Accurate Simulations of Intrinsically Disordered Proteins.Int J Mol Sci. 2020 Aug 26;21(17):6166. doi: 10.3390/ijms21176166. Int J Mol Sci. 2020. PMID: 32859072 Free PMC article.
-
Force field development and simulations of intrinsically disordered proteins.Curr Opin Struct Biol. 2018 Feb;48:40-48. doi: 10.1016/j.sbi.2017.10.008. Epub 2017 Nov 5. Curr Opin Struct Biol. 2018. PMID: 29080468 Free PMC article. Review.
-
Atomic resolution conformational dynamics of intrinsically disordered proteins from NMR spin relaxation.Prog Nucl Magn Reson Spectrosc. 2017 Nov;102-103:43-60. doi: 10.1016/j.pnmrs.2017.06.001. Epub 2017 Jul 10. Prog Nucl Magn Reson Spectrosc. 2017. PMID: 29157493 Review.
Cited by
-
Sequence-Dependent Correlated Segments in the Intrinsically Disordered Region of ChiZ.Biomolecules. 2020 Jun 23;10(6):946. doi: 10.3390/biom10060946. Biomolecules. 2020. PMID: 32585849 Free PMC article.
-
Decreased Water Mobility Contributes To Increased α-Synuclein Aggregation.Angew Chem Weinheim Bergstr Ger. 2023 Feb 6;135(7):e202212063. doi: 10.1002/ange.202212063. Epub 2023 Jan 12. Angew Chem Weinheim Bergstr Ger. 2023. PMID: 38516046 Free PMC article.
-
Molecular simulations integrated with experiments for probing the interaction dynamics and binding mechanisms of intrinsically disordered proteins.Curr Opin Struct Biol. 2024 Feb;84:102756. doi: 10.1016/j.sbi.2023.102756. Epub 2023 Dec 19. Curr Opin Struct Biol. 2024. PMID: 38118365 Free PMC article. Review.
-
Atomistic molecular dynamics simulations of intrinsically disordered proteins.Curr Opin Struct Biol. 2025 Jun;92:103029. doi: 10.1016/j.sbi.2025.103029. Epub 2025 Mar 10. Curr Opin Struct Biol. 2025. PMID: 40068541 Free PMC article. Review.
-
Molecular Basis of Small-Molecule Binding to α-Synuclein.J Am Chem Soc. 2022 Feb 16;144(6):2501-2510. doi: 10.1021/jacs.1c07591. Epub 2022 Feb 7. J Am Chem Soc. 2022. PMID: 35130691 Free PMC article.
References
-
- Levy Y., Onuchic J. N., Water mediation in protein folding and molecular recognition. Annu. Rev. Biophys. Biomol. Struct. 35, 389–415 (2006). - PubMed
-
- Doster W., Cusack S., Petry W., Dynamical transition of myoglobin revealed by inelastic neutron scattering. Nature 337, 754–756 (1989). - PubMed
-
- Rasmussen B. F., Stock A. M., Ringe D., Petsko G. A., Crystalline ribonuclease A loses function below the dynamical transition at 220 K. Nature 357, 423–424 (1992). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

