Sulfation pathways from red to green
- PMID: 31270211
- PMCID: PMC6699852
- DOI: 10.1074/jbc.REV119.007422
Sulfation pathways from red to green
Abstract
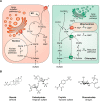
Sulfur is present in the amino acids cysteine and methionine and in a large range of essential coenzymes and cofactors and is therefore essential for all organisms. It is also a constituent of sulfate esters in proteins, carbohydrates, and numerous cellular metabolites. The sulfation and desulfation reactions modifying a variety of different substrates are commonly known as sulfation pathways. Although relatively little is known about the function of most sulfated metabolites, the synthesis of activated sulfate used in sulfation pathways is essential in both animal and plant kingdoms. In humans, mutations in the genes encoding the sulfation pathway enzymes underlie a number of developmental aberrations, and in flies and worms, their loss-of-function is fatal. In plants, a lower capacity for synthesizing activated sulfate for sulfation reactions results in dwarfism, and a complete loss of activated sulfate synthesis is also lethal. Here, we review the similarities and differences in sulfation pathways and associated processes in animals and plants, and we point out how they diverge from bacteria and yeast. We highlight the open questions concerning localization, regulation, and importance of sulfation pathways in both kingdoms and the ways in which findings from these "red" and "green" experimental systems may help reciprocally address questions specific to each of the systems.
Keywords: 3′-phosphoadenosine 5′-phosphosulfate (PAPS) synthase; ATP sulfurylase; Arabidopsis thaliana; adenosine 5′-phosphosulfate kinase; glucosinolates; human; secondary metabolism; steroid hormone; sulfate activation; sulfotransferase; sulfur.
© 2019 Günal et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures


References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

