Cryo_fit: Democratization of flexible fitting for cryo-EM
- PMID: 31279069
- PMCID: PMC7112765
- DOI: 10.1016/j.jsb.2019.05.012
Cryo_fit: Democratization of flexible fitting for cryo-EM
Abstract
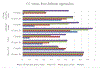
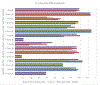
Cryo-electron microscopy (cryo-EM) is becoming a method of choice for describing native conformations of biomolecular complexes at high resolution. The rapid growth of cryo-EM in recent years has created a high demand for automated solutions, both in hardware and software. Flexible fitting of atomic models to three-dimensional (3D) cryo-EM reconstructions by molecular dynamics (MD) simulation is a popular technique but often requires technical expertise in computer simulation. This work introduces cryo_fit, a package for the automatic flexible fitting of atomic models in cryo-EM maps using MD simulation. The package is integrated with the Phenix software suite. The module was designed to automate the multiple steps of MD simulation in a reproducible manner, as well as facilitate refinement and validation through Phenix. Through the use of cryo_fit, scientists with little experience in MD simulation can produce high quality atomic models automatically and better exploit the potential of cryo-EM.
Keywords: Correlation coefficient; Cryo-EM; Flexible fitting; Molecular dynamics; Phenix.
Copyright © 2019. Published by Elsevier Inc.
Figures
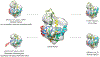
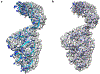

Similar articles
-
NMMD: Efficient Cryo-EM Flexible Fitting Based on Simultaneous Normal Mode and Molecular Dynamics atomic displacements.J Mol Biol. 2022 Apr 15;434(7):167483. doi: 10.1016/j.jmb.2022.167483. Epub 2022 Feb 9. J Mol Biol. 2022. PMID: 35150654
-
MDSPACE: Extracting Continuous Conformational Landscapes from Cryo-EM Single Particle Datasets Using 3D-to-2D Flexible Fitting based on Molecular Dynamics Simulation.J Mol Biol. 2023 May 1;435(9):167951. doi: 10.1016/j.jmb.2023.167951. Epub 2023 Jan 10. J Mol Biol. 2023. PMID: 36638910
-
New tools for the analysis and validation of cryo-EM maps and atomic models.Acta Crystallogr D Struct Biol. 2018 Sep 1;74(Pt 9):814-840. doi: 10.1107/S2059798318009324. Epub 2018 Sep 3. Acta Crystallogr D Struct Biol. 2018. PMID: 30198894 Free PMC article.
-
Current approaches for the fitting and refinement of atomic models into cryo-EM maps using CCP-EM.Acta Crystallogr D Struct Biol. 2018 Jun 1;74(Pt 6):492-505. doi: 10.1107/S2059798318007313. Epub 2018 May 30. Acta Crystallogr D Struct Biol. 2018. PMID: 29872001 Free PMC article. Review.
-
Refinement of Atomic Structures Against cryo-EM Maps.Methods Enzymol. 2016;579:277-305. doi: 10.1016/bs.mie.2016.05.033. Epub 2016 Jun 24. Methods Enzymol. 2016. PMID: 27572731 Review.
Cited by
-
Structural basis for the inhibition of cGAS by nucleosomes.Science. 2020 Oct 23;370(6515):455-458. doi: 10.1126/science.abd0237. Epub 2020 Sep 10. Science. 2020. PMID: 32912999 Free PMC article.
-
Dissemination of pathogenic bacteria is reinforced by a MARTX toxin effector duet.Nat Commun. 2024 Jul 23;15(1):6218. doi: 10.1038/s41467-024-50650-0. Nat Commun. 2024. PMID: 39043696 Free PMC article.
-
Integrative structural studies of the SARS-CoV-2 spike protein during the fusion process (2022).Curr Res Struct Biol. 2022;4:220-230. doi: 10.1016/j.crstbi.2022.06.004. Epub 2022 Jun 23. Curr Res Struct Biol. 2022. PMID: 35765663 Free PMC article.
-
Cryo-EM structure of native human uromodulin, a zona pellucida module polymer.EMBO J. 2020 Dec 15;39(24):e106807. doi: 10.15252/embj.2020106807. Epub 2020 Nov 16. EMBO J. 2020. PMID: 33196145 Free PMC article.
-
Structural investigation of an RNA device that regulates PD-1 expression in mammalian cells.Nucleic Acids Res. 2025 Feb 27;53(5):gkaf156. doi: 10.1093/nar/gkaf156. Nucleic Acids Res. 2025. PMID: 40071935 Free PMC article.
References
-
- Method of the Year 2015. Nat. Methods 13, 1 (2015). - PubMed
-
- Shen PS The 2017 Nobel Prize in Chemistry: cryo-EM comes of age. Anal. Bioanal. Chem 410, 2053–2057 (2018). - PubMed
-
- Danev R & Baumeister W Expanding the boundaries of cryo-EM with phase plates. Curr. Opin. Struct. Biol 46, 87–94 (2017). - PubMed
-
- Murata K & Wolf M Cryo-electron microscopy for structural analysis of dynamic biological macromolecules. Biochim. Biophys. Acta - Gen. Subj 1862, 324–334 (2018). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

