PLAGL2 and POFUT1 are regulated by an evolutionarily conserved bidirectional promoter and are collaboratively involved in colorectal cancer by maintaining stemness
- PMID: 31279780
- PMCID: PMC6642334
- DOI: 10.1016/j.ebiom.2019.06.051
PLAGL2 and POFUT1 are regulated by an evolutionarily conserved bidirectional promoter and are collaboratively involved in colorectal cancer by maintaining stemness
Abstract
Background: Our previous study revealed that PLAGL2 or POFUT1 can promote tumorigenesis and maintain significant positive correlations in colorectal cancer (CRC). However, the mechanism leading to the co-expression and the underlying functional and biological implications remain unclear.
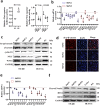
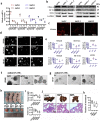
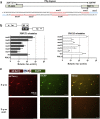
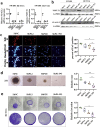
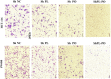
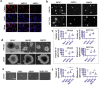
Methods: Clinical tumor tissues and TCGA dataset were utilized to analyze the co-expression of PLAGL2 and POFUT1. Luciferase reporter assays, specially made bidirectional promoter vectors and ectopic expression of 3'UTR were employed to study the mechanisms of co-expression. In vitro and in vivo assays were performed to further confirm the oncogenic function of both. The sphere formation assay, immunofluorescence, Western blot and qRT-PCR were performed to investigate the effect of both genes in colorectal cancer stem cells (CSCs).
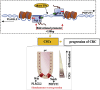
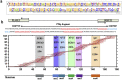
Findings: PLAGL2 and POFUT1 maintained co-expression in CRC (r = 0.91, p < .0001). An evolutionarily conserved bidirectional promoter, rather than post-transcriptional regulation by competing endogenous RNAs, caused the co-expression of PLAGL2 and POFUT1 in CRC. The bidirectional gene pair PLAGL2/POFUT1 was subverted in CRC and acted synergistically to promote colorectal tumorigenesis by maintaining stemness of colorectal cancer stem cells through the Wnt and Notch pathways. Finally, PLAGL2 and POFUT1 share transcription factor binding sites, and introducing mutations into promoter regions with shared transcription regulatory elements led to a decrease in the PLAGL2/POFUT1 promoter activity in both directions.
Interpretation: Our team identified for the first time a bidirectional promoter pair oncogene, PLAGL2-POFUT1, in CRC. The two genes synergistically promote the progression of CRC and affect the characteristics of CSCs, which can offer promising intervention targets for clinicians and researchers. FUND: National Nature Science Foundation of China, the Hunan province projects of Postgraduate Independent Exploration and Innovation of Central South University.
Keywords: Bidirectional promoter; Cancer stem cell; Co-expression; Colorectal cancer; PLAGL2; POFUT1.
Copyright © 2019 The Authors. Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors declare no conflicts of interest that pertain to this work.
Figures





Similar articles
-
POFUT1 and PLAGL2 gene pair linked by a bidirectional promoter: the two in one of tumour progression in colorectal cancer?EBioMedicine. 2019 Aug;46:25-26. doi: 10.1016/j.ebiom.2019.07.065. Epub 2019 Aug 1. EBioMedicine. 2019. PMID: 31378697 Free PMC article. No abstract available.
-
Overexpressed PLAGL2 transcriptionally activates Wnt6 and promotes cancer development in colorectal cancer.Oncol Rep. 2019 Feb;41(2):875-884. doi: 10.3892/or.2018.6914. Epub 2018 Dec 7. Oncol Rep. 2019. PMID: 30535429 Free PMC article.
-
Weighted gene coexpression analysis indicates that PLAGL2 and POFUT1 are related to the differential features of proximal and distal colorectal cancer.Oncol Rep. 2019 Dec;42(6):2473-2485. doi: 10.3892/or.2019.7368. Epub 2019 Oct 14. Oncol Rep. 2019. PMID: 31638246 Free PMC article.
-
Non-coding RNAs Functioning in Colorectal Cancer Stem Cells.Adv Exp Med Biol. 2016;937:93-108. doi: 10.1007/978-3-319-42059-2_5. Adv Exp Med Biol. 2016. PMID: 27573896 Review.
-
Regulatory mechanisms of microRNAs in colorectal cancer and colorectal cancer stem cells.J Cell Physiol. 2020 Feb;235(2):776-789. doi: 10.1002/jcp.29042. Epub 2019 Jul 1. J Cell Physiol. 2020. PMID: 31264216 Review.
Cited by
-
Identifying the tumor location-associated candidate genes in development of new drugs for colorectal cancer using machine-learning-based approach.Med Biol Eng Comput. 2022 Oct;60(10):2877-2897. doi: 10.1007/s11517-022-02641-w. Epub 2022 Aug 10. Med Biol Eng Comput. 2022. PMID: 35948841
-
Pan-Cancer Analysis of Head-to-Head Gene Pairs in Terms of Transcriptional Activity, Co-expression and Regulation.Front Genet. 2021 Jan 7;11:560997. doi: 10.3389/fgene.2020.560997. eCollection 2020. Front Genet. 2021. PMID: 33488665 Free PMC article.
-
Unexplored regulatory sequences of divergently paired GLA and HNRNPH2 loci pertinent to Fabry disease in human kidney and skin cells: Presence of an active bidirectional promoter.Exp Ther Med. 2021 Feb;21(2):154. doi: 10.3892/etm.2020.9586. Epub 2020 Dec 17. Exp Ther Med. 2021. PMID: 33456521 Free PMC article.
-
Identification of potential key genes in gastric cancer using bioinformatics analysis.Biomed Rep. 2020 Apr;12(4):178-192. doi: 10.3892/br.2020.1281. Epub 2020 Feb 20. Biomed Rep. 2020. PMID: 32190306 Free PMC article.
-
Co-regulation and function of FOXM1/RHNO1 bidirectional genes in cancer.Elife. 2021 Apr 23;10:e55070. doi: 10.7554/eLife.55070. Elife. 2021. PMID: 33890574 Free PMC article.
References
-
- Muleris M., Salmon R.J., Dutrillaux A.M. Characteristic chromosomal imbalances in 18 near-diploid colorectal tumors. Cancer Genet Cytogenet. 1987;29:289–301. - PubMed
-
- Carvalho B., Postma C., Mongera S. Multiple putative oncogenes at the chromosome 20q amplicon contribute to colorectal adenoma to carcinoma progression. Gut. 2009;58:79–89. - PubMed
-
- Sillars-Hardebol A.H., Carvalho B., Tijssen M. TPX2 and AURKA promote 20q amplicon-driven colorectal adenoma to carcinoma progression. Gut. 2012;61:1568–1575. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

