Deletion of OGG1 Results in a Differential Signature of Oxidized Purine Base Damage in mtDNA Regions
- PMID: 31284385
- PMCID: PMC6651574
- DOI: 10.3390/ijms20133302
Deletion of OGG1 Results in a Differential Signature of Oxidized Purine Base Damage in mtDNA Regions
Abstract
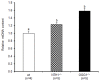
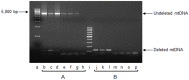
Mitochondrial oxidative stress accumulates with aging and age-related diseases and induces alterations in mitochondrial DNA (mtDNA) content. Since mtDNA qualitative alterations are also associated with aging, repair of mtDNA damage is of great importance. The most relevant form of DNA repair in this context is base excision repair (BER), which removes oxidized bases such as 8-oxoguanine (8-oxoG) and thymine glycol through the action of the mitochondrial isoform of the specific 8-oxoG DNA glycosylase/apurinic or apyrimidinic (AP) lyase (OGG1) or the endonuclease III homolog (NTH1). Mouse strains lacking OGG1 (OGG1-/-) or NTH1 (NTH1-/-) were analyzed for mtDNA alterations. Interestingly, both knockout strains presented a significant increase in mtDNA content, suggestive of a compensatory mtDNA replication. The mtDNA "common deletion" was not detected in either knockout mouse strain, likely because of the young age of the mice. Formamidopyrimidine DNA glycosylase (Fpg)-sensitive sites accumulated in mtDNA from OGG1-/- but not from NTH1-/- mice. Interestingly, the D-loop region was most severely affected by the absence of OGG1, suggesting that this region may be a hotspot for oxidative damage. Thus, we speculate that mtDNA alterations may send a stress message to evoke cell changes through a retrograde mitochondrial-nucleus communication.
Keywords: 8-oxoG localization; NTH1 ko; OGG1 knockout (ko); mtDNA content; mtDNA repair.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Mitochondrial 8-oxoguanine glycosylase decreases mitochondrial fragmentation and improves mitochondrial function in H9C2 cells under oxidative stress conditions.Am J Physiol Cell Physiol. 2014 Feb 1;306(3):C221-9. doi: 10.1152/ajpcell.00140.2013. Epub 2013 Dec 4. Am J Physiol Cell Physiol. 2014. PMID: 24304833 Free PMC article.
-
Targeted deletion of the genes encoding NTH1 and NEIL1 DNA N-glycosylases reveals the existence of novel carcinogenic oxidative damage to DNA.DNA Repair (Amst). 2009 Jul 4;8(7):786-94. doi: 10.1016/j.dnarep.2009.03.001. Epub 2009 Apr 5. DNA Repair (Amst). 2009. PMID: 19346169 Free PMC article.
-
The basal levels of 8-oxoG and other oxidative modifications in intact mitochondrial DNA are low even in repair-deficient (Ogg1(-/-)/Csb(-/-)) mice.Mutat Res. 2007 Dec 1;625(1-2):155-63. doi: 10.1016/j.mrfmmm.2007.06.006. Epub 2007 Jun 27. Mutat Res. 2007. PMID: 17675188
-
Repair of 8-oxo-7,8-dihydroguanine in prokaryotic and eukaryotic cells: Properties and biological roles of the Fpg and OGG1 DNA N-glycosylases.Free Radic Biol Med. 2017 Jun;107:179-201. doi: 10.1016/j.freeradbiomed.2016.11.042. Epub 2016 Nov 27. Free Radic Biol Med. 2017. PMID: 27903453 Review.
-
Multiple DNA glycosylases for repair of 8-oxoguanine and their potential in vivo functions.Prog Nucleic Acid Res Mol Biol. 2001;68:193-205. doi: 10.1016/s0079-6603(01)68100-5. Prog Nucleic Acid Res Mol Biol. 2001. PMID: 11554297 Review.
Cited by
-
The Age-Sensitive Efficacy of Calorie Restriction on Mitochondrial Biogenesis and mtDNA Damage in Rat Liver.Int J Mol Sci. 2021 Feb 7;22(4):1665. doi: 10.3390/ijms22041665. Int J Mol Sci. 2021. PMID: 33562258 Free PMC article.
-
Small-Molecule Inhibitor of 8-Oxoguanine DNA Glycosylase 1 Regulates Inflammatory Responses during Pseudomonas aeruginosa Infection.J Immunol. 2020 Oct 15;205(8):2231-2242. doi: 10.4049/jimmunol.1901533. Epub 2020 Sep 14. J Immunol. 2020. PMID: 32929043 Free PMC article.
-
mtDNA and Mitochondrial Stress Signaling in Human Diseases: A Special Issue.Int J Mol Sci. 2020 Apr 9;21(7):2617. doi: 10.3390/ijms21072617. Int J Mol Sci. 2020. PMID: 32283804 Free PMC article.
-
Heritable pattern of oxidized DNA base repair coincides with pre-targeting of repair complexes to open chromatin.Nucleic Acids Res. 2021 Jan 11;49(1):221-243. doi: 10.1093/nar/gkaa1120. Nucleic Acids Res. 2021. PMID: 33300026 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

