Widespread Translational Control of Fibrosis in the Human Heart by RNA-Binding Proteins
- PMID: 31284728
- PMCID: PMC6749977
- DOI: 10.1161/CIRCULATIONAHA.119.039596
Widespread Translational Control of Fibrosis in the Human Heart by RNA-Binding Proteins
Abstract
Background: Fibrosis is a common pathology in many cardiac disorders and is driven by the activation of resident fibroblasts. The global posttranscriptional mechanisms underlying fibroblast-to-myofibroblast conversion in the heart have not been explored.
Methods: Genome-wide changes of RNA transcription and translation during human cardiac fibroblast activation were monitored with RNA sequencing and ribosome profiling. We then used RNA-binding protein-based analyses to identify translational regulators of fibrogenic genes. The integration with cardiac ribosome occupancy levels of 30 dilated cardiomyopathy patients demonstrates that these posttranscriptional mechanisms are also active in the diseased fibrotic human heart.
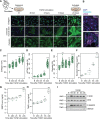
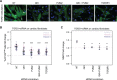
Results: We generated nucleotide-resolution translatome data during the transforming growth factor β1-driven cellular transition of human cardiac fibroblasts to myofibroblasts. This identified dynamic changes of RNA transcription and translation at several time points during the fibrotic response, revealing transient and early-responder genes. Remarkably, about one-third of all changes in gene expression in activated fibroblasts are subject to translational regulation, and dynamic variation in ribosome occupancy affects protein abundance independent of RNA levels. Targets of RNA-binding proteins were strongly enriched in posttranscriptionally regulated genes, suggesting genes such as MBNL2 can act as translational activators or repressors. Ribosome occupancy in the hearts of patients with dilated cardiomyopathy suggested the same posttranscriptional regulatory network was underlying cardiac fibrosis. Key network hubs include RNA-binding proteins such as Pumilio RNA binding family member 2 (PUM2) and Quaking (QKI) that work in concert to regulate the translation of target transcripts in human diseased hearts. Furthermore, silencing of both PUM2 and QKI inhibits the transition of fibroblasts toward profibrotic myofibroblasts in response to transforming growth factor β1.
Conclusions: We reveal widespread translational effects of transforming growth factor β1 and define novel posttranscriptional regulatory networks that control the fibroblast-to-myofibroblast transition. These networks are active in human heart disease, and silencing of hub genes limits fibroblast activation. Our findings show the central importance of translational control in fibrosis and highlight novel pathogenic mechanisms in heart failure.
Keywords: RNA-binding proteins; TGF-beta1; dilated cardiomyopathy; fibrosis; ribosome profiling.
Figures





Comment in
-
Cardiac fibrosis research: two steps forward.Nat Rev Cardiol. 2019 Sep;16(9):515. doi: 10.1038/s41569-019-0245-7. Nat Rev Cardiol. 2019. PMID: 31341287 No abstract available.
References
-
- Rockey DC, Bell PD, Hill JA. Fibrosis–a common pathway to organ injury and failure. N Engl J Med. 2015;372:1138–1149. doi: 10.1056/NEJMra1300575. - PubMed
-
- Burstein B, Nattel S. Atrial fibrosis: mechanisms and clinical relevance in atrial fibrillation. J Am Coll Cardiol. 2008;51:802–809. doi: 10.1016/j.jacc.2007.09.064. - PubMed
-
- Teekakirikul P, Eminaga S, Toka O, Alcalai R, Wang L, Wakimoto H, Nayor M, Konno T, Gorham JM, Wolf CM, et al. Cardiac fibrosis in mice with hypertrophic cardiomyopathy is mediated by non-myocyte proliferation and requires Tgf-β. J Clin Invest. 2010;120:3520–3529. doi: 10.1172/JCI42028. - PMC - PubMed
-
- Gulati A, Jabbour A, Ismail TF, Guha K, Khwaja J, Raza S, Morarji K, Brown TD, Ismail NA, Dweck MR, et al. Association of fibrosis with mortality and sudden cardiac death in patients with nonischemic dilated cardiomyopathy. JAMA. 2013;309:896–908. doi: 10.1001/jama.2013.1363. - PubMed
-
- Schelbert EB, Fridman Y, Wong TC, Abu Daya H, Piehler KM, Kadakkal A, Miller CA, Ugander M, Maanja M, Kellman P, et al. Temporal relation between myocardial fibrosis and heart failure with preserved ejection fraction: association with baseline disease severity and subsequent outcome. JAMA Cardiol. 2017;2:995–1006. doi: 10.1001/jamacardio.2017.2511. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

