DNA hypermethylation in disease: mechanisms and clinical relevance
- PMID: 31284823
- PMCID: PMC6791695
- DOI: 10.1080/15592294.2019.1638701
DNA hypermethylation in disease: mechanisms and clinical relevance
Abstract
Increasing numbers of studies implicate abnormal DNA methylation in cancer and many non-malignant diseases. This is consistent with numerous findings about differentiation-associated changes in DNA methylation at promoters, enhancers, gene bodies, and sites that control higher-order chromatin structure. Abnormal increases or decreases in DNA methylation contribute to or are markers for cancer formation and tumour progression. Aberrant DNA methylation is also associated with neurological diseases, immunological diseases, atherosclerosis, and osteoporosis. In this review, I discuss DNA hypermethylation in disease and its interrelationships with normal development as well as proposed mechanisms for the origin of and pathogenic consequences of disease-associated hypermethylation. Disease-linked DNA hypermethylation can help drive oncogenesis partly by its effects on cancer stem cells and by the CpG island methylator phenotype (CIMP); atherosclerosis by disease-related cell transdifferentiation; autoimmune and neurological diseases through abnormal perturbations of cell memory; and diverse age-associated diseases by age-related accumulation of epigenetic alterations.
Keywords: CpG island methylator phenotype (CIMP); DNA hypermethylation; DNMT; TET; aging; atherosclerosis; brain disease; cancer stem cells; immune dysfunction; osteoporosis.
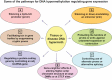
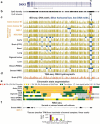
Figures
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
