Structure of the eukaryotic protein O-mannosyltransferase Pmt1-Pmt2 complex
- PMID: 31285605
- PMCID: PMC6684406
- DOI: 10.1038/s41594-019-0262-6
Structure of the eukaryotic protein O-mannosyltransferase Pmt1-Pmt2 complex
Abstract
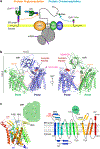
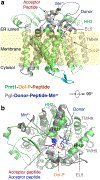
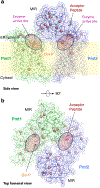
In eukaryotes, a nascent peptide entering the endoplasmic reticulum (ER) is scanned by two Sec61 translocon-associated large membrane machines for protein N-glycosylation and protein O-mannosylation, respectively. While the structure of the eight-protein oligosaccharyltransferase complex has been determined recently, the structures of mannosyltransferases of the PMT family, which are an integral part of ER protein homeostasis, are still unknown. Here we report cryo-EM structures of the Saccharomyces cerevisiae Pmt1-Pmt2 complex bound to a donor and an acceptor peptide at 3.2-Å resolution, showing that each subunit contains 11 transmembrane helices and a lumenal β-trefoil fold termed the MIR domain. The structures reveal the substrate recognition model and confirm an inverting mannosyl-transferring reaction mechanism by the enzyme complex. Furthermore, we found that the transmembrane domains of Pmt1 and Pmt2 share a structural fold with the catalytic subunits of oligosaccharyltransferases, confirming a previously proposed evolutionary relationship between protein O-mannosylation and protein N-glycosylation.
Conflict of interest statement
Competing Interests
The authors declare no competing financial interests.
Figures


References
-
- VanderVen BC, Harder JD, Crick DC & Belisle JT Export-mediated assembly of mycobacterial glycoproteins parallels eukaryotic pathways. Science 309, 941–3 (2005). - PubMed
-
- Xu C, Wang S, Thibault G & Ng DT Futile protein folding cycles in the ER are terminated by the unfolded protein O-mannosylation pathway. Science 340, 978–81 (2013). - PubMed
-
- Loibl M & Strahl S Protein O-mannosylation: what we have learned from baker’s yeast. Biochim Biophys Acta 1833, 2438–46 (2013). - PubMed
-
- Burda P & Aebi M The dolichol pathway of N-linked glycosylation. Biochim Biophys Acta 1426, 239–57 (1999). - PubMed
Methods-only References
-
- Otwinowski Z & Minor W Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol 276, 307–326 (1997). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

