Identification of microRNA-92a and the related combination biomarkers as promising substrates in predicting risk, recurrence and poor survival of colorectal cancer
- PMID: 31289586
- PMCID: PMC6603388
- DOI: 10.7150/jca.30306
Identification of microRNA-92a and the related combination biomarkers as promising substrates in predicting risk, recurrence and poor survival of colorectal cancer
Abstract
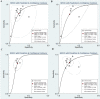
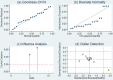
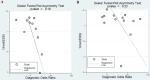
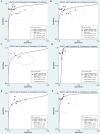
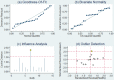
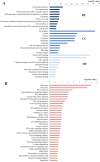
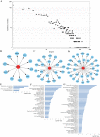
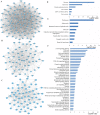
Background: Previous studies demonstrated that microRNA-92a (miR-92a) may serve as a novel promising biomarker in colorectal cancer (CRC) patients. However, a comprehensive analysis of the contribution of miR-92a in CRC is lacking. We aimed to systematically summarize the diagnostic and prognostic values of miR-92a in CRC. Methods: The diagnostic and prognostic roles of individual miR-92a and the combination biomarkers based on miR-92a were evaluated through comprehensive meta-analyses. Meanwhile, the function and potential mechanisms of miR-92a were assessed by an integrative bioinformatics analysis. Results: According to the results, we found that miR-92a yielded a pooled area under ROC curve (AUC) of 0.82 (sensitivity: 76%, specificity: 75%) in discriminating CRC from controls. Notably, the combination biomarkers based on miR-92a increased the diagnostic performance, yielding an AUC of 0.91, with a sensitivity of 83% and a specificity of 87%. For the prognostic meta-analysis, patients with higher expression of miR-92a had significant shorter overall survival (pooled HR: 2.30; 95% CI: 1.03-5.12). In addition, the regulated genes of miR-92a were retrieved and enriched through gene ontology and pathway analysis, indicating their correlations with the initiation and progression of CRC. Furthermore, protein-protein interaction network was set up with miR-92a targets and screened for hub nodes and significant modules, which were confirmed strongly involved in the occurrence and development of CRC again. Conclusions: Current evidences suggest miR-92a is a promising biomarker for early detection and prognosis of CRC while miRNA combination biomarkers may be considered as the right way for clinical practice. However, more prospective studies are required to highlight the theoretical strengths.
Keywords: Biomarker; Colorectal cancer; Meta-analysis; System biological analysis.
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures

Similar articles
-
Identification of microRNA-181 as a promising biomarker for predicting the poor survival in colorectal cancer.Cancer Med. 2019 Oct;8(13):5995-6009. doi: 10.1002/cam4.2520. Epub 2019 Aug 25. Cancer Med. 2019. PMID: 31448575 Free PMC article.
-
Integrated analyses of microRNA-29 family and the related combination biomarkers demonstrate their widespread influence on risk, recurrence, metastasis and survival outcome in colorectal cancer.Cancer Cell Int. 2019 Jul 15;19:181. doi: 10.1186/s12935-019-0907-x. eCollection 2019. Cancer Cell Int. 2019. PMID: 31346316 Free PMC article.
-
Evaluation of microRNA 92a Expression and Its Target Protein Bim in Colorectal Cancer.Asian Pac J Cancer Prev. 2022 Feb 1;23(2):723-730. doi: 10.31557/APJCP.2022.23.2.723. Asian Pac J Cancer Prev. 2022. PMID: 35225486 Free PMC article.
-
Biomarker roles identification of miR-106 family for predicting the risk and poor survival of colorectal cancer.BMC Cancer. 2020 Jun 3;20(1):506. doi: 10.1186/s12885-020-06863-9. BMC Cancer. 2020. PMID: 32493303 Free PMC article.
-
The Efficacy of miR-20a as a Diagnostic and Prognostic Biomarker for Colorectal Cancer: A Systematic Review and Meta-Analysis.Cancers (Basel). 2019 Aug 3;11(8):1111. doi: 10.3390/cancers11081111. Cancers (Basel). 2019. PMID: 31382594 Free PMC article. Review.
Cited by
-
Identification of microRNA-181 as a promising biomarker for predicting the poor survival in colorectal cancer.Cancer Med. 2019 Oct;8(13):5995-6009. doi: 10.1002/cam4.2520. Epub 2019 Aug 25. Cancer Med. 2019. PMID: 31448575 Free PMC article.
-
Comprehensive evaluation of microRNA-10b in digestive system cancers reveals prognostic implication and signaling pathways associated with tumor progression.J Cancer. 2021 May 13;12(13):4011-4024. doi: 10.7150/jca.51303. eCollection 2021. J Cancer. 2021. PMID: 34093806 Free PMC article.
-
Knockdown of miR-92a suppresses the stemness of colorectal cancer cells via mediating SOCS3.Bioengineered. 2022 Mar;13(3):5613-5624. doi: 10.1080/21655979.2021.2022267. Bioengineered. 2022. PMID: 35184640 Free PMC article.
-
Gut Microbiota-MicroRNA Interactions in Intestinal Homeostasis and Cancer Development.Microorganisms. 2022 Dec 31;11(1):107. doi: 10.3390/microorganisms11010107. Microorganisms. 2022. PMID: 36677399 Free PMC article. Review.
-
CDH1 overexpression predicts bladder cancer from early stage and inversely correlates with immune infiltration.BMC Urol. 2022 Sep 21;22(1):156. doi: 10.1186/s12894-022-01103-7. BMC Urol. 2022. PMID: 36131343 Free PMC article.
References
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30. - PubMed
-
- Siegel RL, Miller KD, Fedewa SA, Ahnen DJ, Meester RGS, Barzi A. et al. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67:177–93. - PubMed
-
- Schreuders EH, Ruco A, Rabeneck L, Schoen RE, Sung JJ, Young GP. et al. Colorectal cancer screening: a global overview of existing programmes. Gut. 2015;64:1637–49. - PubMed
-
- Robertson DJ, Imperiale TF. Stool Testing for Colorectal Cancer Screening. Gastroenterology. 2015;149:1286–93. - PubMed
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97. - PubMed
LinkOut - more resources
Full Text Sources

