De novo sequencing of tree peony (Paeonia suffruticosa) transcriptome to identify critical genes involved in flowering and floral organ development
- PMID: 31296170
- PMCID: PMC6624964
- DOI: 10.1186/s12864-019-5857-0
De novo sequencing of tree peony (Paeonia suffruticosa) transcriptome to identify critical genes involved in flowering and floral organ development
Abstract
Background: Tree peony (Paeonia suffruticosa Andrews) is a globally famous ornamental flower, with large and colorful flowers and abundant flower types. However, a relatively short and uniform flowering period hinders the applications and production of ornamental tree peony. Unfortunately, the molecular mechanism of regulating flowering time and floral organ development in tree peony has yet to be elucidated. Because of the absence of genomic information, 454-based transcriptome sequence technology for de novo transcriptomics was used to identify the critical flowering genes using re-blooming, non-re-blooming, and wild species of tree peonies.
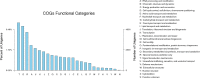
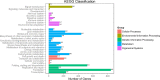
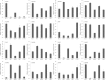
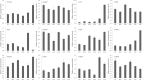
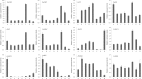
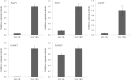
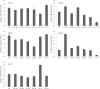
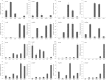
Results: A total of 29,275 unigenes were obtained from the bud transcriptome, with an N50 of 776 bp. The average length of unigenes was 677.18 bp, and the longest sequence was 5815 bp. Functional annotation showed that 22,823, 17,321, 13,312, 20,041, and 9940 unigenes were annotated by NCBI-NR, Swiss-Prot, COG, GO, and KEGG, respectively. Within the differentially expressed genes (DEGs) 64 flowering-related genes were identified and some important flowering genes were also characterized by bioinformatics methods, reverse transcript polymerase chain reaction (RT-PCR), and rapid-amplification of cDNA ends (RACE). Then, the putative genetic network of flowering induction pathways and a floral organ development model were put forward, according to the comparisons of DEGs in any two samples and expression levels of the important flowering genes in differentiated buds, buds from different developmental stages, and with GA or vernalization treated. In tree peony, five pathways (long day, vernalization, autonomous, age, and gibberellin) regulated flowering, and the floral organ development followed an ABCE model. Moreover, it was also found that the genes PsAP1, PsCOL1, PsCRY1, PsCRY2, PsFT, PsLFY, PsLHY, PsGI, PsSOC1, and PsVIN3 probably regulated re-blooming of tree peony.
Conclusion: This study provides a comprehensive report on the flowering-related genes in tree peony for the first time and investigated the expression levels of the critical flowering related genes in buds of different cultivars, developmental stages, differentiated primordium, and flower parts. These results could provide valuable insights into the molecular mechanisms of flowering time regulation and floral organ development.
Keywords: Floral model; Flowering induction pathway; MADS-box gene; Re-blooming; Transcriptome; Tree peony.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





References
-
- Wang SL, Xue JQ, Ahmadi N, Holloway P, Zhu FY, Ren XX, Zhang XX. Molecular characterization and expression patterns of PsSVP genes reveal distinct roles in flower bud abortion and flowering in tree peony (Paeonia Suffruticosa) Can J Plant Sci. 2014;94:1181–1193. doi: 10.4141/cjps2013-360. - DOI
MeSH terms
Grants and funding
- 31501800/National Natural Science Foundation of China
- 31572156/National Natural Science Foundation of China
- 2015QRNC001/China Association for Science and Technology Foundation for Young Scholars (CN)
- CARS-21/China Agriculture Research System
- CAAS-ASTIPIVFCAAS/the Agricultural Science and Technology Innovation Program (ASTIP) of the Chinese Academy of Agricultural Sciences
LinkOut - more resources
Full Text Sources

