Reconstructing Tree-Child Networks from Reticulate-Edge-Deleted Subnetworks
- PMID: 31297691
- PMCID: PMC6764941
- DOI: 10.1007/s11538-019-00641-w
Reconstructing Tree-Child Networks from Reticulate-Edge-Deleted Subnetworks
Abstract
Network reconstruction lies at the heart of phylogenetic research. Two well-studied classes of phylogenetic networks include tree-child networks and level-k networks. In a tree-child network, every non-leaf node has a child that is a tree node or a leaf. In a level-k network, the maximum number of reticulations contained in a biconnected component is k. Here, we show that level-k tree-child networks are encoded by their reticulate-edge-deleted subnetworks, which are subnetworks obtained by deleting a single reticulation edge, if [Formula: see text]. Following this, we provide a polynomial-time algorithm for uniquely reconstructing such networks from their reticulate-edge-deleted subnetworks. Moreover, we show that this can even be done when considering subnetworks obtained by deleting one reticulation edge from each biconnected component with k reticulations.
Keywords: Network encoding; Phylogenetic network; Reticulate-edge-deleted subnetworks; Tree-child networks.
Figures
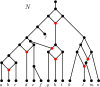
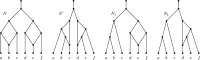
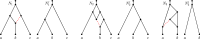
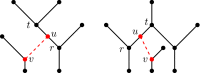
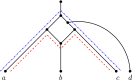
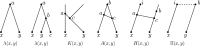
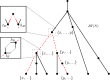
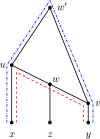
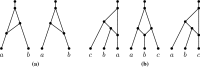
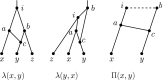
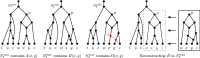

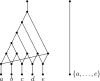
Similar articles
-
Embedding gene trees into phylogenetic networks by conflict resolution algorithms.Algorithms Mol Biol. 2022 May 19;17(1):11. doi: 10.1186/s13015-022-00218-8. Algorithms Mol Biol. 2022. PMID: 35590416 Free PMC article.
-
When is a Phylogenetic Network Simply an Amalgamation of Two Trees?Bull Math Biol. 2018 Sep;80(9):2338-2348. doi: 10.1007/s11538-018-0463-x. Epub 2018 Jul 6. Bull Math Biol. 2018. PMID: 29981001
-
Computing the Bounds of the Number of Reticulations in a Tree-Child Network That Displays a Set of Trees.J Comput Biol. 2024 Apr;31(4):345-359. doi: 10.1089/cmb.2023.0309. Epub 2024 Jan 29. J Comput Biol. 2024. PMID: 38285528
-
Not all phylogenetic networks are leaf-reconstructible.J Math Biol. 2019 Oct;79(5):1623-1638. doi: 10.1007/s00285-019-01405-9. Epub 2019 Jul 30. J Math Biol. 2019. PMID: 31363828 Free PMC article.
-
Classes of explicit phylogenetic networks and their biological and mathematical significance.J Math Biol. 2022 May 3;84(6):47. doi: 10.1007/s00285-022-01746-y. J Math Biol. 2022. PMID: 35503141 Review.
Cited by
-
Embedding gene trees into phylogenetic networks by conflict resolution algorithms.Algorithms Mol Biol. 2022 May 19;17(1):11. doi: 10.1186/s13015-022-00218-8. Algorithms Mol Biol. 2022. PMID: 35590416 Free PMC article.
-
Clustering systems of phylogenetic networks.Theory Biosci. 2023 Nov;142(4):301-358. doi: 10.1007/s12064-023-00398-w. Epub 2023 Aug 12. Theory Biosci. 2023. PMID: 37573261 Free PMC article.
-
The tree of blobs of a species network: identifiability under the coalescent.J Math Biol. 2022 Dec 6;86(1):10. doi: 10.1007/s00285-022-01838-9. J Math Biol. 2022. PMID: 36472708 Free PMC article.
-
Applicability of several rooted phylogenetic network algorithms for representing the evolutionary history of SARS-CoV-2.BMC Ecol Evol. 2021 Dec 7;21(1):220. doi: 10.1186/s12862-021-01946-y. BMC Ecol Evol. 2021. PMID: 34876022 Free PMC article.
References
-
- Bordewich M, Semple C, Tokac N. Constructing tree-child networks from distance matrices. Algorithmica. 2018;80(8):2240–2259. doi: 10.1007/s00453-017-0320-6. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

