Spectrin-based membrane skeleton supports ciliogenesis
- PMID: 31299042
- PMCID: PMC6655744
- DOI: 10.1371/journal.pbio.3000369
Spectrin-based membrane skeleton supports ciliogenesis
Abstract
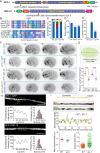
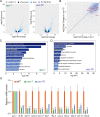
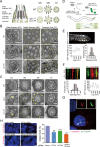
Cilia are remarkable cellular devices that power cell motility and transduce extracellular signals. To assemble a cilium, a cylindrical array of 9 doublet microtubules push out an extension of the plasma membrane. Membrane tension regulates cilium formation; however, molecular pathways that link mechanical stimuli to ciliogenesis are unclear. Using genome editing, we introduced hereditary elliptocytosis (HE)- and spinocerebellar ataxia (SCA)-associated mutations into the Caenorhabditis elegans membrane skeletal protein spectrin. We show that these mutations impair mechanical support for the plasma membrane and change cell shape. RNA sequencing (RNA-seq) analyses of spectrin-mutant animals uncovered a global down-regulation of ciliary gene expression, prompting us to investigate whether spectrin participates in ciliogenesis. Spectrin mutations affect intraflagellar transport (IFT), disrupt axonemal microtubules, and inhibit cilium formation, and the endogenous spectrin periodically distributes along cilia. Mammalian spectrin also localizes in cilia and regulates ciliogenesis. These results define a previously unrecognized yet conserved role of spectrin-based mechanical support for cilium biogenesis.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
Whole-Organism Developmental Expression Profiling Identifies RAB-28 as a Novel Ciliary GTPase Associated with the BBSome and Intraflagellar Transport.PLoS Genet. 2016 Dec 8;12(12):e1006469. doi: 10.1371/journal.pgen.1006469. eCollection 2016 Dec. PLoS Genet. 2016. PMID: 27930654 Free PMC article.
-
Optimal sidestepping of intraflagellar transport kinesins regulates structure and function of sensory cilia.EMBO J. 2020 Jun 17;39(12):e103955. doi: 10.15252/embj.2019103955. Epub 2020 Apr 27. EMBO J. 2020. PMID: 32338401 Free PMC article.
-
Sensory ciliogenesis in Caenorhabditis elegans: assignment of IFT components into distinct modules based on transport and phenotypic profiles.Mol Biol Cell. 2007 May;18(5):1554-69. doi: 10.1091/mbc.e06-09-0805. Epub 2007 Feb 21. Mol Biol Cell. 2007. PMID: 17314406 Free PMC article.
-
Ciliogenesis: building the cell's antenna.Nat Rev Mol Cell Biol. 2011 Apr;12(4):222-34. doi: 10.1038/nrm3085. Nat Rev Mol Cell Biol. 2011. PMID: 21427764 Review.
-
Ciliary diffusion barrier: the gatekeeper for the primary cilium compartment.Cytoskeleton (Hoboken). 2011 Jun;68(6):313-24. doi: 10.1002/cm.20514. Epub 2011 Jun 10. Cytoskeleton (Hoboken). 2011. PMID: 21634025 Free PMC article. Review.
Cited by
-
The neuron-intrinsic membrane skeleton is required for motor neuron integrity throughout lifespan.bioRxiv [Preprint]. 2025 Feb 24:2025.02.23.639536. doi: 10.1101/2025.02.23.639536. bioRxiv. 2025. PMID: 40060495 Free PMC article. Preprint.
-
Plastin and spectrin cooperate to stabilize the actomyosin cortex during cytokinesis.Curr Biol. 2021 Dec 20;31(24):5415-5428.e10. doi: 10.1016/j.cub.2021.09.055. Epub 2021 Oct 18. Curr Biol. 2021. PMID: 34666005 Free PMC article.
-
A kinesin-1 adaptor complex controls bimodal slow axonal transport of spectrin in Caenorhabditis elegans.Dev Cell. 2023 Oct 9;58(19):1847-1863.e12. doi: 10.1016/j.devcel.2023.08.031. Epub 2023 Sep 25. Dev Cell. 2023. PMID: 37751746 Free PMC article.
-
Actin filament debranching regulates cell polarity during cell migration and asymmetric cell division.Proc Natl Acad Sci U S A. 2021 Sep 14;118(37):e2100805118. doi: 10.1073/pnas.2100805118. Proc Natl Acad Sci U S A. 2021. PMID: 34507987 Free PMC article.
-
An interphase pool of KIF11 localizes at the basal bodies of primary cilia and a reduction in KIF11 expression alters cilia dynamics.Sci Rep. 2020 Aug 18;10(1):13946. doi: 10.1038/s41598-020-70787-4. Sci Rep. 2020. PMID: 32811879 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

