TREM2 Acts Downstream of CD33 in Modulating Microglial Pathology in Alzheimer's Disease
- PMID: 31301936
- PMCID: PMC6728215
- DOI: 10.1016/j.neuron.2019.06.010
TREM2 Acts Downstream of CD33 in Modulating Microglial Pathology in Alzheimer's Disease
Abstract
The microglial receptors CD33 and TREM2 have been associated with risk for Alzheimer's disease (AD). Here, we investigated crosstalk between CD33 and TREM2. We showed that knockout of CD33 attenuated amyloid beta (Aβ) pathology and improved cognition in 5xFAD mice, both of which were abrogated by additional TREM2 knockout. Knocking out TREM2 in 5xFAD mice exacerbated Aβ pathology and neurodegeneration but reduced Iba1+ cell numbers, all of which could not be rescued by additional CD33 knockout. RNA-seq profiling of microglia revealed that genes related to phagocytosis and signaling (IL-6, IL-8, acute phase response) are upregulated in 5xFAD;CD33-/- and downregulated in 5xFAD;TREM2-/- mice. Differential gene expression in 5xFAD;CD33-/- microglia depended on the presence of TREM2, suggesting TREM2 acts downstream of CD33. Crosstalk between CD33 and TREM2 includes regulation of the IL-1β/IL-1RN axis and a gene set in the "receptor activity chemokine" cluster. Our results should facilitate AD therapeutics targeting these receptors.
Keywords: Alzheimer's; CD33; IL-1beta; RNA-seq; TREM2; amyloid beta; microglia; neuroinflammation; pathway analysis; transcriptomics.
Copyright © 2019 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interest
The authors declare no competing interests.
Figures



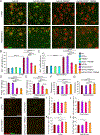



Comment in
-
Bringing Order out of Chaos: Establishing an Epistatic Relationship between CD33 and TREM2.Neuron. 2019 Sep 4;103(5):747-749. doi: 10.1016/j.neuron.2019.08.019. Neuron. 2019. PMID: 31487521 Free PMC article.
References
-
- Ben-Menachem-Zidon O, Ben-Menahem Y, Ben-Hur T, and Yirmiya R (2014). Intra-hippocampal transplantation of neural precursor cells with transgenic over-expression of IL-1 receptor antagonist rescues memory and neurogenesis impairments in an Alzheimer’s disease model. Neuropsychopharmacology 39, 401–414. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

