Mechanisms of Progression of Myeloid Preleukemia to Transformed Myeloid Leukemia in Children with Down Syndrome
- PMID: 31303423
- PMCID: PMC6863161
- DOI: 10.1016/j.ccell.2019.06.007
Mechanisms of Progression of Myeloid Preleukemia to Transformed Myeloid Leukemia in Children with Down Syndrome
Erratum in
-
Mechanisms of Progression of Myeloid Preleukemia to Transformed Myeloid Leukemia in Children with Down Syndrome.Cancer Cell. 2019 Sep 16;36(3):340. doi: 10.1016/j.ccell.2019.08.014. Cancer Cell. 2019. PMID: 31526763 No abstract available.
Abstract
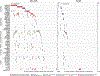
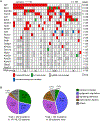
Myeloid leukemia in Down syndrome (ML-DS) clonally evolves from transient abnormal myelopoiesis (TAM), a preleukemic condition in DS newborns. To define mechanisms of leukemic transformation, we combined exome and targeted resequencing of 111 TAM and 141 ML-DS samples with functional analyses. TAM requires trisomy 21 and truncating mutations in GATA1; additional TAM variants are usually not pathogenic. By contrast, in ML-DS, clonal and subclonal variants are functionally required. We identified a recurrent and oncogenic hotspot gain-of-function mutation in myeloid cytokine receptor CSF2RB. By a multiplex CRISPR/Cas9 screen in an in vivo murine TAM model, we tested loss-of-function of 22 recurrently mutated ML-DS genes. Loss of 18 different genes produced leukemias that phenotypically, genetically, and transcriptionally mirrored ML-DS.
Keywords: Acute myeloid leukemia; CRISPR screen; Down syndrome; GATA1; cancer transformation; preleukemia.
Copyright © 2019 Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
D.R. has consulting/advisory roles for Roche, Celgene, Hexal, Pfizer, Novartis, and Boehringer, and receives Celgene research funding. D.R. received travel, accommodation, and expenses from Jazz Pharmaceuticals and Griffols. J.D.C. receives research funding from Scholar Rock and Forma Therapeutics. All other authors declare no competing interests.
Figures




Comment in
-
Mechanisms of Leukemia Evolution: Lessons from a Congenital Syndrome.Cancer Cell. 2019 Aug 12;36(2):115-117. doi: 10.1016/j.ccell.2019.07.004. Cancer Cell. 2019. PMID: 31408616
References
-
- Ahmed M, Sternberg A, Hall G, Thomas A, Smith O, O’Marcaigh A, Wynn R, Stevens R, Addison M, King D, et al. (2004). Natural history of GATA1 mutations in Down syndrome. Blood 103, 2480–2489. - PubMed
-
- Alford KA, Reinhardt K, Garnett C, Norton A, Bohmer K, von Neuhoff C, Kolenova A, Marchi E, Klusmann JH, Roberts I, et al. (2011). Analysis of GATA1 mutations in Down syndrome transient myeloproliferative disorder and myeloid leukemia. Blood 118, 2222–2238. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

