Role of the msaABCR Operon in Cell Wall Biosynthesis, Autolysis, Integrity, and Antibiotic Resistance in Staphylococcus aureus
- PMID: 31307991
- PMCID: PMC6761503
- DOI: 10.1128/AAC.00680-19
Role of the msaABCR Operon in Cell Wall Biosynthesis, Autolysis, Integrity, and Antibiotic Resistance in Staphylococcus aureus
Abstract
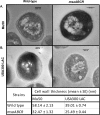
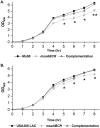
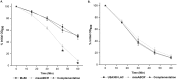
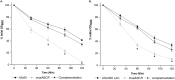
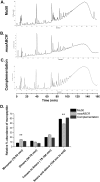
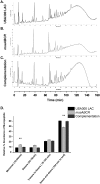
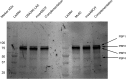
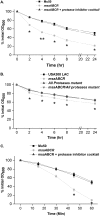
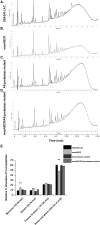
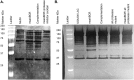
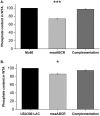
Staphylococcus aureus is an important human pathogen in both community and health care settings. One of the challenges with S. aureus as a pathogen is its acquisition of antibiotic resistance. Previously, we showed that deletion of the msaABCR operon reduces cell wall thickness, resulting in decreased resistance to vancomycin in vancomycin-intermediate S. aureus (VISA). In this study, we investigated the nature of the cell wall defect in the msaABCR operon mutant in the Mu50 (VISA) and USA300 LAC methicillin-resistant Staphylococcus aureus (MRSA) strains. Results showed that msaABCR mutant cells had decreased cross-linking in both strains. This defect is typically due to increased murein hydrolase activity and/or nonspecific processing of murein hydrolases mediated by increased protease activity in mutant cells. The defect was enhanced by a decrease in teichoic acid content in the msaABCR mutant. Therefore, we propose that deletion of the msaABCR operon results in decreased peptidoglycan cross-linking, leading to increased susceptibility toward cell wall-targeting antibiotics, such as β-lactams and vancomycin. Moreover, we also observed significantly downregulated transcription of early cell wall-synthesizing genes, supporting the finding that msaABCR mutant cells have decreased peptidoglycan synthesis. More specifically, the msaABCR mutant in the USA300 LAC strain (MRSA) showed significantly reduced expression of the murA gene, whereas the msaABCR mutant in the Mu50 strain (VISA) showed significantly reduced expression of glmU, murA, and murD Thus, we conclude that the msaABCR operon controls the balance between cell wall synthesis and cell wall hydrolysis, which is required for maintaining a robust cell wall and acquiring resistance to cell wall-targeting antibiotics, such as vancomycin and the β-lactams.
Keywords: Staphylococcus aureus; antibiotic resistance; cell wall; msaABCR; peptidoglycan.
Copyright © 2019 American Society for Microbiology.
Figures


Similar articles
-
Complete Reconstitution of the Vancomycin-Intermediate Staphylococcus aureus Phenotype of Strain Mu50 in Vancomycin-Susceptible S. aureus.Antimicrob Agents Chemother. 2016 May 23;60(6):3730-42. doi: 10.1128/AAC.00420-16. Print 2016 Jun. Antimicrob Agents Chemother. 2016. PMID: 27067329 Free PMC article.
-
The msaABCR operon regulates resistance in vancomycin-intermediate Staphylococcus aureus strains.Antimicrob Agents Chemother. 2014 Nov;58(11):6685-95. doi: 10.1128/AAC.03280-14. Epub 2014 Aug 25. Antimicrob Agents Chemother. 2014. PMID: 25155591 Free PMC article.
-
The role of the msaABCR operon in implant-associated chronic osteomyelitis in Staphylococcus aureus USA300 LAC.BMC Microbiol. 2020 Oct 27;20(1):324. doi: 10.1186/s12866-020-01964-8. BMC Microbiol. 2020. PMID: 33109085 Free PMC article.
-
Staphylococcus aureus response and adaptation to vancomycin.Adv Microb Physiol. 2024;85:201-258. doi: 10.1016/bs.ampbs.2024.04.006. Epub 2024 Jun 1. Adv Microb Physiol. 2024. PMID: 39059821 Review.
-
Mechanisms of vancomycin resistance in Staphylococcus aureus.J Clin Invest. 2014 Jul;124(7):2836-40. doi: 10.1172/JCI68834. Epub 2014 Jul 1. J Clin Invest. 2014. PMID: 24983424 Free PMC article. Review.
Cited by
-
Celastrol mitigates staphyloxanthin biosynthesis and biofilm formation in Staphylococcus aureus via targeting key regulators of virulence; in vitro and in vivo approach.BMC Microbiol. 2022 Apr 15;22(1):106. doi: 10.1186/s12866-022-02515-z. BMC Microbiol. 2022. PMID: 35421933 Free PMC article.
-
Staphylococcus aureus Biofilm-Associated Infections: Have We Found a Clinically Relevant Target?Microorganisms. 2025 Apr 9;13(4):852. doi: 10.3390/microorganisms13040852. Microorganisms. 2025. PMID: 40284688 Free PMC article. Review.
-
Antibiofilm and staphyloxanthin inhibitory potential of terbinafine against Staphylococcus aureus: in vitro and in vivo studies.Ann Clin Microbiol Antimicrob. 2022 May 30;21(1):21. doi: 10.1186/s12941-022-00513-7. Ann Clin Microbiol Antimicrob. 2022. PMID: 35637481 Free PMC article.
-
The Role of β-Glycosylated Wall Teichoic Acids in the Reduction of Vancomycin Susceptibility in Vancomycin-Intermediate Staphylococcus aureus.Microbiol Spectr. 2021 Oct 31;9(2):e0052821. doi: 10.1128/Spectrum.00528-21. Epub 2021 Oct 20. Microbiol Spectr. 2021. PMID: 34668723 Free PMC article.
-
Molecular Mechanisms and Epidemiology of Fosfomycin Resistance in Staphylococcus aureus Isolated From Patients at a Teaching Hospital in China.Front Microbiol. 2020 Jun 26;11:1290. doi: 10.3389/fmicb.2020.01290. eCollection 2020. Front Microbiol. 2020. PMID: 32670230 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

