PRDM9 forms a trimer by interactions within the zinc finger array
- PMID: 31308055
- PMCID: PMC6643046
- DOI: 10.26508/lsa.201800291
PRDM9 forms a trimer by interactions within the zinc finger array
Abstract
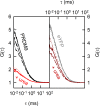
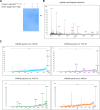
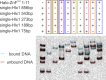
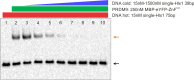
PRDM9 is a trans-acting factor directing meiotic recombination to specific DNA-binding sites by its zinc finger (ZnF) array. It was suggested that PRDM9 is a multimer; however, we do not know the stoichiometry or the components inducing PRDM9 multimerization. In this work, we used in vitro binding studies and characterized with electrophoretic mobility shift assays, mass spectrometry, and fluorescence correlation spectroscopy the stoichiometry of the PRDM9 multimer of two different murine PRDM9 alleles carrying different tags and domains produced with different expression systems. Based on the migration distance of the PRDM9-DNA complex, we show that PRDM9 forms a trimer. Moreover, this stoichiometry is adapted already by the free, soluble protein with little exchange between protein monomers. The variable ZnF array of PRDM9 is sufficient for multimerization, and at least five ZnFs form already a functional trimer. Finally, we also show that only one ZnF array within the PRDM9 oligomer binds to the DNA, whereas the remaining two ZnF arrays likely maintain the trimer by ZnF-ZnF interactions.
© 2019 Schwarz et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

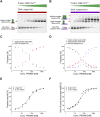

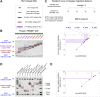
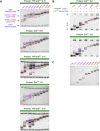

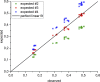



Similar articles
-
The long zinc finger domain of PRDM9 forms a highly stable and long-lived complex with its DNA recognition sequence.Chromosome Res. 2017 Jun;25(2):155-172. doi: 10.1007/s10577-017-9552-1. Epub 2017 Feb 2. Chromosome Res. 2017. PMID: 28155083 Free PMC article.
-
A map of human PRDM9 binding provides evidence for novel behaviors of PRDM9 and other zinc-finger proteins in meiosis.Elife. 2017 Oct 26;6:e28383. doi: 10.7554/eLife.28383. Elife. 2017. PMID: 29072575 Free PMC article.
-
Diversity of Prdm9 zinc finger array in wild mice unravels new facets of the evolutionary turnover of this coding minisatellite.PLoS One. 2014 Jan 13;9(1):e85021. doi: 10.1371/journal.pone.0085021. eCollection 2014. PLoS One. 2014. PMID: 24454780 Free PMC article.
-
The case of the fickle fingers: how the PRDM9 zinc finger protein specifies meiotic recombination hotspots in humans.PLoS Biol. 2011 Dec;9(12):e1001211. doi: 10.1371/journal.pbio.1001211. Epub 2011 Dec 6. PLoS Biol. 2011. PMID: 22162947 Free PMC article. Review.
-
The consequences of sequence erosion in the evolution of recombination hotspots.Philos Trans R Soc Lond B Biol Sci. 2017 Dec 19;372(1736):20160462. doi: 10.1098/rstb.2016.0462. Philos Trans R Soc Lond B Biol Sci. 2017. PMID: 29109225 Free PMC article. Review.
Cited by
-
A deleterious frameshift insertion mutation in the ZNF142 gene leads to intellectual developmental disorder with impaired speech in three affected siblings: Clinical features and literature review.Mol Genet Genomic Med. 2023 Dec;11(12):e2261. doi: 10.1002/mgg3.2261. Epub 2023 Jul 26. Mol Genet Genomic Med. 2023. PMID: 37496384 Free PMC article. Review.
-
Multiple Genomic Landscapes of Recombination and Genomic Divergence in Wild Populations of House Mice-The Role of Chromosomal Fusions and Prdm9.Mol Biol Evol. 2024 Apr 2;41(4):msae063. doi: 10.1093/molbev/msae063. Mol Biol Evol. 2024. PMID: 38513632 Free PMC article.
-
PRDM9 drives the location and rapid evolution of recombination hotspots in salmonid fish.PLoS Biol. 2025 Jan 6;23(1):e3002950. doi: 10.1371/journal.pbio.3002950. eCollection 2025 Jan. PLoS Biol. 2025. PMID: 39761307 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
