Cellular response to moderate chromatin architectural defects promotes longevity
- PMID: 31309140
- PMCID: PMC6620092
- DOI: 10.1126/sciadv.aav1165
Cellular response to moderate chromatin architectural defects promotes longevity
Abstract
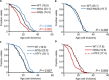
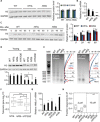
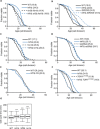
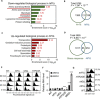
Changes in chromatin organization occur during aging. Overexpression of histones partially alleviates these changes and promotes longevity. We report that deletion of the histone H3-H4 minor locus HHT1-HHF1 extended the replicative life span of Saccharomyces cerevisiae. This longevity effect was mediated through TOR signaling inhibition. We present evidence for evolutionarily conserved transcriptional and phenotypic responses to defects in chromatin structure, collectively termed the chromatin architectural defect (CAD) response. Promoters of the CAD response genes were sensitive to histone dosage, with HHT1-HHF1 deletion, nucleosome occupancy was reduced at these promoters allowing transcriptional activation induced by stress response transcription factors Msn2 and Gis1, both of which were required for the life-span extension of hht1-hhf1Δ. Therefore, we conclude that the CAD response induced by moderate chromatin defects promotes longevity.
Figures
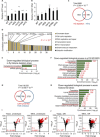

Similar articles
-
Elevated histone expression promotes life span extension.Mol Cell. 2010 Sep 10;39(5):724-35. doi: 10.1016/j.molcel.2010.08.015. Mol Cell. 2010. PMID: 20832724 Free PMC article.
-
Loss of Nat4 and its associated histone H4 N-terminal acetylation mediates calorie restriction-induced longevity.EMBO Rep. 2016 Dec;17(12):1829-1843. doi: 10.15252/embr.201642540. Epub 2016 Oct 31. EMBO Rep. 2016. PMID: 27799288 Free PMC article.
-
The retrograde response links metabolism with stress responses, chromatin-dependent gene activation, and genome stability in yeast aging.Gene. 2005 Jul 18;354:22-7. doi: 10.1016/j.gene.2005.03.040. Gene. 2005. PMID: 15890475
-
Histone post-translational modifications regulate transcription and silent chromatin in Saccharomyces cerevisiae.Ernst Schering Res Found Workshop. 2006;(57):127-53. doi: 10.1007/3-540-37633-x_8. Ernst Schering Res Found Workshop. 2006. PMID: 16568953 Review.
-
Histone variants: key players of chromatin.Cell Tissue Res. 2014 Jun;356(3):457-66. doi: 10.1007/s00441-014-1862-4. Epub 2014 Apr 30. Cell Tissue Res. 2014. PMID: 24781148 Review.
Cited by
-
Loss of chromatin structural integrity is a source of stress during aging.Hum Genet. 2020 Mar;139(3):371-380. doi: 10.1007/s00439-019-02100-x. Epub 2020 Jan 3. Hum Genet. 2020. PMID: 31900586 Free PMC article. Review.
-
Unraveling Histone Loss in Aging and Senescence.Cells. 2024 Feb 9;13(4):320. doi: 10.3390/cells13040320. Cells. 2024. PMID: 38391933 Free PMC article. Review.
-
A mortality timer based on nucleolar size triggers nucleolar integrity loss and catastrophic genomic instability.Nat Aging. 2024 Dec;4(12):1782-1793. doi: 10.1038/s43587-024-00754-5. Epub 2024 Nov 25. Nat Aging. 2024. PMID: 39587368 Free PMC article.
-
Decidualization of human endometrial stromal cells requires steroid receptor coactivator-3.Front Reprod Health. 2022 Nov 24;4:1033581. doi: 10.3389/frph.2022.1033581. eCollection 2022. Front Reprod Health. 2022. PMID: 36505394 Free PMC article.
-
The flavonoid corylin exhibits lifespan extension properties in mouse.Nat Commun. 2022 Mar 9;13(1):1238. doi: 10.1038/s41467-022-28908-2. Nat Commun. 2022. PMID: 35264584 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

