Spotting, Transcription and In Situ Synthesis: Three Routes for the Fabrication of RNA Microarrays
- PMID: 31321002
- PMCID: PMC6612525
- DOI: 10.1016/j.csbj.2019.06.004
Spotting, Transcription and In Situ Synthesis: Three Routes for the Fabrication of RNA Microarrays
Abstract
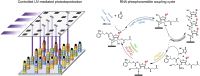
DNA microarrays have become commonplace in the last two decades, but the synthesis of other nucleic acids biochips, most importantly RNA, has only recently been developed to a similar extent. RNA microarrays can be seen as organized surfaces displaying a potentially very large number of unique sequences and are of invaluable help in understanding the complexity of RNA structure and function as they allow the probing and treatment of each of the many different sequences simultaneously. Three approaches have emerged for the fabrication of RNA microarrays. The earliest examples used a direct, manual or mechanical, deposition of pre-synthesized, purified RNA oligonucleotides onto the surface in a process called spotting. In a second approach, pre-spotted or in situ-synthesized DNA microarrays are employed as templates for the transcription of RNA, subsequently or immediately captured on the surface. Finally, a third approach attempts to mirror the phosphoramidite-based protocols for in situ synthesis of high-density DNA arrays in order to produce in situ synthesized RNA microarrays. In this mini-review, we describe the chemistry and the engineering behind the fabrications methods, underlining the advantages and shortcomings of each, and illustrate how versatile these platforms can be by presenting some of their applications.
Keywords: Microarray; Phosphoramidites; Photolithography; RNA; Solid-phase synthesis; Spotting; Transcription; in situ RNA synthesis.
Figures


References
-
- Sharp P.A. The centrality of RNA. Cell. 2009;136:577–580. - PubMed
-
- Wilson T.J., Liu Y.J., Lilley D.M.J. Ribozymes and the mechanisms that underlie RNA catalysis. Front Chem Sci Eng. 2016;10:178–185.
-
- Hentze M.W., Castello A., Schwarzl T., Preiss T. A brave new world of RNA-binding proteins. Nat Rev Mol Cell Biol. 2018;19:327–341. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
