Insights into the Potential of the Atlantic Cod Gut Microbiome as Biomarker of Oil Contamination in the Marine Environment
- PMID: 31336609
- PMCID: PMC6680985
- DOI: 10.3390/microorganisms7070209
Insights into the Potential of the Atlantic Cod Gut Microbiome as Biomarker of Oil Contamination in the Marine Environment
Abstract
Background: Microorganisms are widespread in all environments, including in and on animal bodies. The gut microbiome has an essential influence on fish health, and is affected by several persistent and harmful organic and inorganic contaminants. Considering the shifts in gut microbiota composition observed in those studies, we hypothesized that certain microbial groups in the gut can serve as indicators of pollution. To test this hypothesis, we explored the possibility of identifying key microbial players that indicate environmental contamination.
Methods: Published 16S rRNA gene amplicon sequencing data generated from the gut microbiota of Atlantic cod caught in geographically different Norwegian waters were used for bacterial diversity comparison.
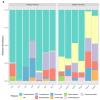
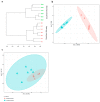
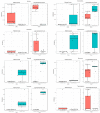
Results: Different microbiomes were identified between the northern Norway and southern Norway samples. Several bacterial genera previously identified as polycyclic aromatic hydrocarbon degraders were present only in the samples collected in the southern Norway area, suggesting fish contamination with oil-related compounds.
Conclusions: The results contribute to the identification of bacterial taxa present in the Atlantic cod gut that indicate fish exposure to contaminants in the marine environment.
Keywords: animal; gut; marine; microbiome; prokaryote.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures


Similar articles
-
Closely-related Photobacterium strains comprise the majority of bacteria in the gut of migrating Atlantic cod (Gadus morhua).Microbiome. 2019 Apr 17;7(1):64. doi: 10.1186/s40168-019-0681-y. Microbiome. 2019. PMID: 30995938 Free PMC article.
-
A Single Vibrionales 16S rRNA Oligotype Dominates the Intestinal Microbiome in Two Geographically Separated Atlantic cod Populations.Front Microbiol. 2018 Jul 13;9:1561. doi: 10.3389/fmicb.2018.01561. eCollection 2018. Front Microbiol. 2018. PMID: 30057577 Free PMC article.
-
Interpopulation Variation in the Atlantic Salmon Microbiome Reflects Environmental and Genetic Diversity.Appl Environ Microbiol. 2018 Aug 1;84(16):e00691-18. doi: 10.1128/AEM.00691-18. Print 2018 Aug 15. Appl Environ Microbiol. 2018. PMID: 29915104 Free PMC article.
-
Gut Microbiomes of the Eastern Oyster (Crassostrea virginica) and the Blue Mussel (Mytilus edulis): Temporal Variation and the Influence of Marine Aggregate-Associated Microbial Communities.mSphere. 2019 Dec 11;4(6):e00730-19. doi: 10.1128/mSphere.00730-19. mSphere. 2019. PMID: 31826972 Free PMC article.
-
Divergent bacterial landscapes: unraveling geographically driven microbiomes in Atlantic cod.Sci Rep. 2024 Mar 13;14(1):6088. doi: 10.1038/s41598-024-56616-y. Sci Rep. 2024. PMID: 38480867 Free PMC article.
Cited by
-
The nasal microbiota of two marine fish species: diversity, community structure, variability, and first insights into the impacts of climate change-related stressors.FEMS Microbiol Ecol. 2025 Feb 20;101(3):fiaf018. doi: 10.1093/femsec/fiaf018. FEMS Microbiol Ecol. 2025. PMID: 39963731 Free PMC article.
-
Reveal your microbes, and i'll reveal your origins: geographical traceability via Scomber colias intestinal tract metagenomics.Anim Microbiome. 2025 May 6;7(1):43. doi: 10.1186/s42523-025-00398-9. Anim Microbiome. 2025. PMID: 40329323 Free PMC article.
-
Hatchery tanks induce intense reduction in microbiota diversity associated with gills and guts of two endemic species of the São Francisco River.Front Microbiol. 2022 Dec 2;13:966436. doi: 10.3389/fmicb.2022.966436. eCollection 2022. Front Microbiol. 2022. PMID: 36532494 Free PMC article.
-
Do fish gut microbiotas vary across spatial scales? A case study of Diplodus vulgaris in the Mediterranean Sea.Anim Microbiome. 2024 Jun 13;6(1):32. doi: 10.1186/s42523-024-00319-2. Anim Microbiome. 2024. PMID: 38872229 Free PMC article.
-
Effects of Different Preservation Methods on the Structure and Diversity of Intestinal Microbiota of Marine Fishes.Curr Microbiol. 2025 Jan 13;82(2):81. doi: 10.1007/s00284-025-04060-0. Curr Microbiol. 2025. PMID: 39804371
References
-
- Martínez-Porchas M., Vargas-Albores F. Microbial metagenomics in aquaculture: A potential tool for a deeper insight into the activity. Rev. Aquac. 2017;9:42–56. doi: 10.1111/raq.12102. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

