Epigenetics, N-myrystoyltransferase-1 and casein kinase-2-alpha modulates the increased replication of HIV-1 CRF02_AG, compared to subtype-B viruses
- PMID: 31337802
- PMCID: PMC6650493
- DOI: 10.1038/s41598-019-47069-9
Epigenetics, N-myrystoyltransferase-1 and casein kinase-2-alpha modulates the increased replication of HIV-1 CRF02_AG, compared to subtype-B viruses
Abstract
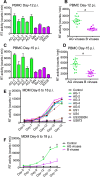
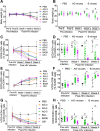
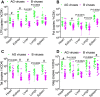
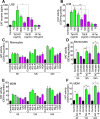
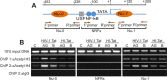
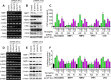
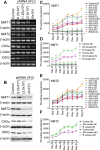
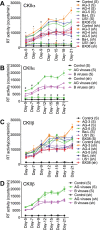
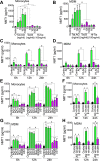
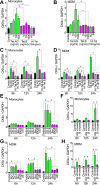
HIV subtypes distribution varies by geographic regions; this is likely associated with differences in viral fitness but the predictors and underlying mechanisms are unknown. Using in-vitro, in-vivo, and ex-vivo approaches, we found significantly higher transactivation and replication of HIV-1-CRF02_AG (prevalent throughout West-Central Africa), compared to subtype-B. While CRF02_AG-infected animals showed higher viremia, subtype-B-infected animals showed significantly more weight loss, lower CD4+ T-cells and lower CD4/CD8 ratios, suggesting that factors other than viremia contribute to immunosuppression and wasting syndrome in HIV/AIDS. Compared to HIV-1-subtype-B and its Tat proteins(Tat.B), HIV-1-CRF02_AG and Tat.AG significantly increased histone acetyl-transferase activity and promoter histones H3 and H4 acetylation. Silencing N-myrystoyltransferase(NMT)-1 and casein-kinase-(CK)-II-alpha prevented Tat.AG- and HIV-1-CRF02_AG-mediated viral transactivation and replication, but not Tat.B- or HIV-1-subtype-B-mediated effects. Tat.AG and HIV-1-CRF02_AG induced the expression of NMT-1 and CKII-alpha in human monocytes and macrophages, but Tat.B and HIV-1-subtype-B had no effect. These data demonstrate that NMT1, CKII-alpha, histone acetylation and histone acetyl-transferase modulate the increased replication of HIV-1-CRF02_AG. These novel findings demonstrate that HIV genotype influence viral replication and provide insights into the molecular mechanisms of differential HIV-1 replication. These studies underline the importance of considering the influence of viral genotypes in HIV/AIDS epidemiology, replication, and eradication strategies.
Conflict of interest statement
The authors declare no competing interests.
Figures

References
-
- Nijhuis, M., van Maarseveen, N. M. & Boucher, C. A. Antiviral resistance and impact on viral replication capacity: evolution of viruses under antiviral pressure occurs in three phases. Handb Exp Pharmacol, 299–320, 10.1007/978-3-540-79086-0_11 (2009). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

