Integrating Gene and Protein Expression Reveals Perturbed Functional Networks in Alzheimer's Disease
- PMID: 31340147
- PMCID: PMC7503200
- DOI: 10.1016/j.celrep.2019.06.073
Integrating Gene and Protein Expression Reveals Perturbed Functional Networks in Alzheimer's Disease
Abstract
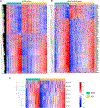
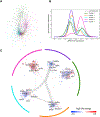
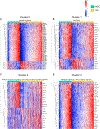
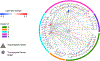
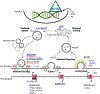
Asymptomatic and symptomatic Alzheimer's disease (AD) subjects may present with equivalent neuropathological burdens but have significantly different antemortem cognitive decline rates. Using the transcriptome as a proxy for functional state, we selected 414 expression profiles of symptomatic AD subjects and age-matched non-demented controls from a community-based neuropathological study. By combining brain tissue-specific protein interactomes with gene networks, we identified functionally distinct composite clusters of genes that reveal extensive changes in expression levels in AD. Global expression for clusters broadly corresponding to synaptic transmission, metabolism, cell cycle, survival, and immune response were downregulated, while the upregulated cluster included largely uncharacterized processes. We propose that loss of EGR3 regulation mediates synaptic deficits by targeting the synaptic vesicle cycle. Our results highlight the utility of integrating protein interactions with gene perturbations to generate a comprehensive framework for characterizing alterations in the molecular network as applied to AD.
Keywords: Alzheimer’s disease; Louvain algorithm; clustering; gene-protein networks; network analysis; protein-protein interaction; transcriptional regulators; transcriptome.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors declare no competing interests.
Figures

Similar articles
-
Integrative network analysis of nineteen brain regions identifies molecular signatures and networks underlying selective regional vulnerability to Alzheimer's disease.Genome Med. 2016 Nov 1;8(1):104. doi: 10.1186/s13073-016-0355-3. Genome Med. 2016. PMID: 27799057 Free PMC article.
-
Transcriptional alterations related to neuropathology and clinical manifestation of Alzheimer's disease.PLoS One. 2012;7(11):e48751. doi: 10.1371/journal.pone.0048751. Epub 2012 Nov 7. PLoS One. 2012. PMID: 23144955 Free PMC article.
-
Cell cycle checkpoint abnormalities during dementia: A plausible association with the loss of protection against oxidative stress in Alzheimer's disease [corrected].PLoS One. 2013 Jul 5;8(7):e68361. doi: 10.1371/journal.pone.0068361. Print 2013. PLoS One. 2013. PMID: 23861893 Free PMC article.
-
Systems biology and gene networks in Alzheimer's disease.Neurosci Biobehav Rev. 2019 Jan;96:31-44. doi: 10.1016/j.neubiorev.2018.11.007. Epub 2018 Nov 20. Neurosci Biobehav Rev. 2019. PMID: 30465785 Review.
-
Cerebrospinal fluid biomarkers of neurovascular dysfunction in mild dementia and Alzheimer's disease.J Cereb Blood Flow Metab. 2015 Jul;35(7):1055-68. doi: 10.1038/jcbfm.2015.76. Epub 2015 Apr 22. J Cereb Blood Flow Metab. 2015. PMID: 25899298 Free PMC article. Review.
Cited by
-
Protein mishandling and impaired lysosomal proteolysis generated through calcium dysregulation in Alzheimer's disease.Proc Natl Acad Sci U S A. 2022 Dec 6;119(49):e2211999119. doi: 10.1073/pnas.2211999119. Epub 2022 Nov 28. Proc Natl Acad Sci U S A. 2022. PMID: 36442130 Free PMC article.
-
RTP801/REDD1 contributes to neuroinflammation severity and memory impairments in Alzheimer's disease.Cell Death Dis. 2021 Jun 15;12(6):616. doi: 10.1038/s41419-021-03899-y. Cell Death Dis. 2021. PMID: 34131105 Free PMC article.
-
A novel human tau knock-in mouse model reveals interaction of Abeta and human tau under progressing cerebral amyloidosis in 5xFAD mice.Alzheimers Res Ther. 2023 Jan 14;15(1):16. doi: 10.1186/s13195-022-01144-y. Alzheimers Res Ther. 2023. PMID: 36641439 Free PMC article.
-
Transcriptional Networks of Microglia in Alzheimer's Disease and Insights into Pathogenesis.Genes (Basel). 2019 Oct 12;10(10):798. doi: 10.3390/genes10100798. Genes (Basel). 2019. PMID: 31614849 Free PMC article. Review.
-
Molecular signatures underlying neurofibrillary tangle susceptibility in Alzheimer's disease.Neuron. 2022 Sep 21;110(18):2929-2948.e8. doi: 10.1016/j.neuron.2022.06.021. Epub 2022 Jul 25. Neuron. 2022. PMID: 35882228 Free PMC article.
References
-
- Alazami AM, Patel N, Shamseldin HE, Anazi S, Al-Dosari MS, Alzahrani F, Hijazi H, Alshammari M, Aldahmesh MA, Salih MA, et al. (2015). Accelerating novel candidate gene discovery in neurogenetic disorders via whole-exome sequencing of prescreened multiplex consanguineous families. Cell Rep. 10, 148–161. - PubMed
-
- Atamna H, and Frey WH 2nd. (2007). Mechanisms of mitochondrial dysfunction and energy deficiency in Alzheimer’s disease. Mitochondrion 7, 297–310. - PubMed

