Deep RNA-Seq reveals miRNome differences in mammary tissue of lactating Holstein and Montbéliarde cows
- PMID: 31362707
- PMCID: PMC6668132
- DOI: 10.1186/s12864-019-5987-4
Deep RNA-Seq reveals miRNome differences in mammary tissue of lactating Holstein and Montbéliarde cows
Abstract
Background: Genetic polymorphisms are known to influence milk production and composition. However, the genomic mechanisms involved in the genetic regulation of milk component synthesis are not completely understood. MicroRNAs (miRNAs) regulate gene expression. Previous research suggests that the high developmental potential of the mammary gland may depend in part on a specific miRNA expression pattern. The objective of the present study was to compare the mammary gland miRNomes of two dairy cow breeds, Holstein and Montbéliarde, which have different mammogenic potentials that are related to differences in dairy performance.
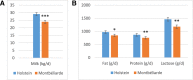
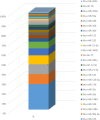
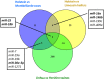
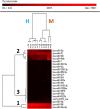
Results: Milk, fat, protein, and lactose yields were lower in Montbéliarde cows than in Holstein cows. We detected 754 distinct miRNAs in the mammary glands of Holstein (n = 5) and Montbéliarde (n = 6) midlactating cows using RNA-Seq technology, among which 738 were known and 16 were predicted miRNAs. The 25 most abundant miRNAs accounted for 90.6% of the total reads. The comparison of their abundances in the mammary glands of Holstein versus Montbéliarde cows identified 22 differentially expressed miRNAs (Padj ≤ 0.05). Among them, 11 presented a fold change ≥2, and 2 (miR-100 and miR-146b) were highly expressed. Among the most abundant miRNAs, miR-186 is known to inhibit cell proliferation and epithelial-to-mesenchymal transition. Data mining showed that 17 differentially expressed miRNAs with more than 20 reads were involved in the regulation of mammary gland plasticity. Several of them may potentially target mRNAs involved in signaling pathways (such as mTOR) and lipid metabolism, thereby indicating that they could influence milk composition.
Conclusion: We found differences in the mammary gland miRNomes of two dairy cattle breeds. These differences suggest a potential role for miRNAs in mammary gland plasticity and milk component synthesis, both of which are related to milk production and composition. Further research is warranted on the genetic regulation of miRNAs and their role in milk synthesis.
Keywords: Breeds; Dairy cows; Differentially expressed; Mammary gland; MiRNome; RNA-Seq.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
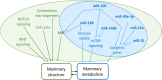
References
-
- Ferlay A, Martin B, Lerch S, Gobert M, Pradel P, Chilliard Y. Effects of supplementation of maize silage diets with extruded linseed, vitamin E and plant extracts rich in polyphenols, and morning v. evening milking on milk fatty acid profiles in Holstein and Montbeliarde cows. Animal. 2010;4(4):627–640. doi: 10.1017/S1751731109991224. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

